Main Analyses
Library
## Lade nötiges Paket: carData## lattice theme set by effectsTheme()
## See ?effectsTheme for details.##
## Attache Paket: 'sjstats'## Die folgenden Objekte sind maskiert von 'package:effectsize':
##
## cohens_f, cramers_v, phi## Lade nötiges Paket: lme4## Lade nötiges Paket: Matrix##
## Attache Paket: 'lmerTest'## Das folgende Objekt ist maskiert 'package:lme4':
##
## lmer## Das folgende Objekt ist maskiert 'package:stats':
##
## step##
## Attache Paket: 'dplyr'## Die folgenden Objekte sind maskiert von 'package:formr':
##
## first, last## Die folgenden Objekte sind maskiert von 'package:stats':
##
## filter, lag## Die folgenden Objekte sind maskiert von 'package:base':
##
## intersect, setdiff, setequal, union## ── Attaching core tidyverse packages ──────────────────────── tidyverse 2.0.0 ──
## ✔ forcats 1.0.0 ✔ stringr 1.5.1
## ✔ lubridate 1.9.4 ✔ tibble 3.2.1
## ✔ purrr 1.0.2 ✔ tidyr 1.3.1
## ✔ readr 2.1.5## ── Conflicts ────────────────────────────────────────── tidyverse_conflicts() ──
## ✖ tidyr::expand() masks Matrix::expand()
## ✖ dplyr::filter() masks stats::filter()
## ✖ dplyr::first() masks formr::first()
## ✖ dplyr::lag() masks stats::lag()
## ✖ dplyr::last() masks formr::last()
## ✖ tidyr::pack() masks Matrix::pack()
## ✖ tidyr::unpack() masks Matrix::unpack()
## ℹ Use the conflicted package (<http://conflicted.r-lib.org/>) to force all conflicts to become errors## Lade nötiges Paket: nlme
##
## Attache Paket: 'nlme'
##
## Das folgende Objekt ist maskiert 'package:dplyr':
##
## collapse
##
## Das folgende Objekt ist maskiert 'package:lme4':
##
## lmList
##
## This is mgcv 1.9-1. For overview type 'help("mgcv-package")'.## Lade nötiges Paket: zoo
##
## Attache Paket: 'zoo'
##
## Die folgenden Objekte sind maskiert von 'package:base':
##
## as.Date, as.Date.numericData
Load selected data based on 03_codebook
Political, Ethnic, and Religious Similarity
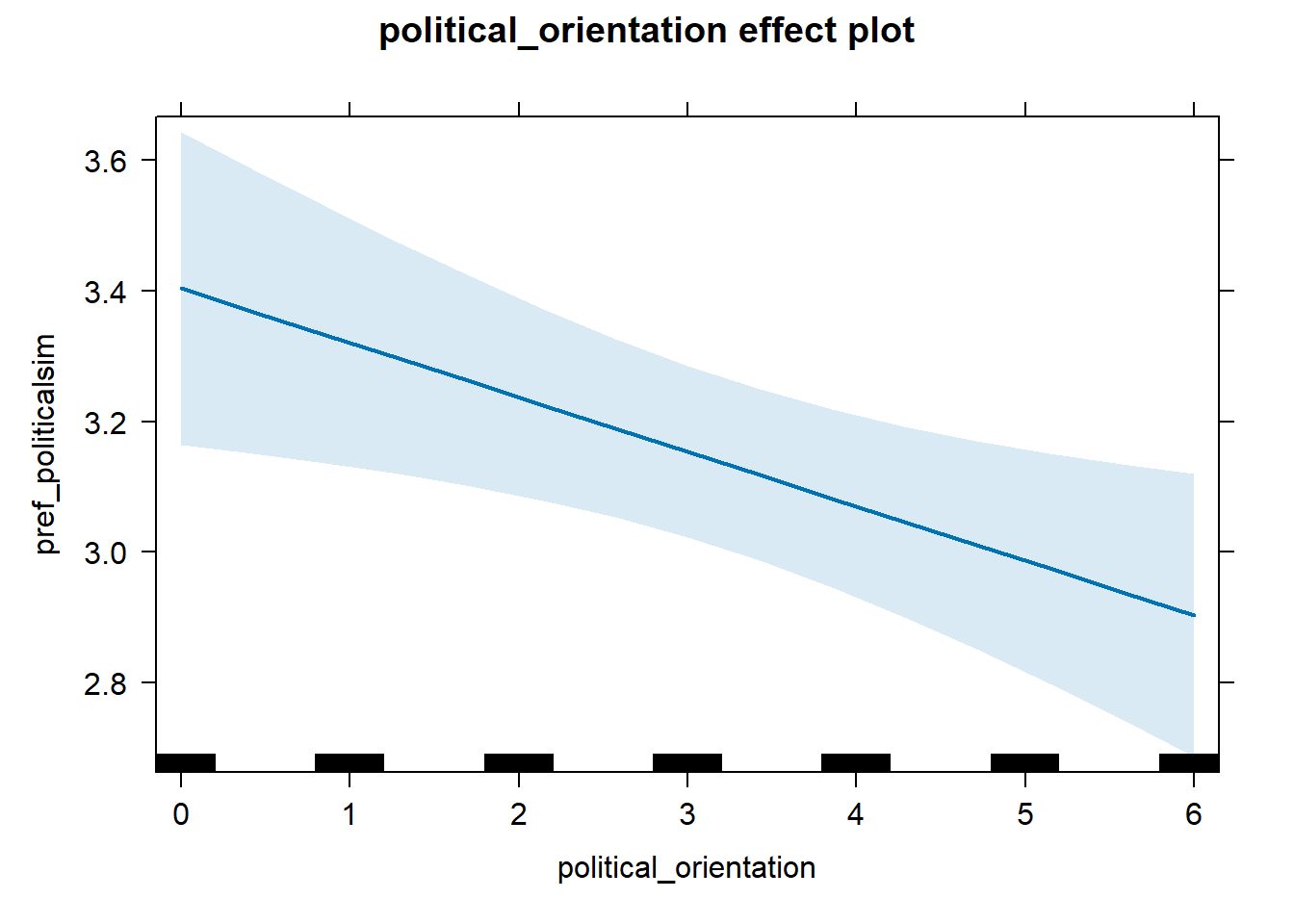
H1a Preference for Similarity in Political Beliefs
H1a(1) There is no linear link between right-wing political orientation and women’s preferences for partner’s similar political beliefs and values. H1a(2) There is a positive quadratic link between right-wing political orientation and women’s preferences for partner’s similar political beliefs and values. Outcome: Preference ratings for partner’s similar political beliefs and values. Predictor: Political Orientation. Random intercept and random slope for country.
H1a(1) Linear Effect
Models
Summary
## Linear mixed model fit by REML. t-tests use Satterthwaite's method [
## lmerModLmerTest]
## Formula:
## pref_politicalsim ~ political_orientation + (1 + political_orientation |
## country)
## Data: data_included_documented
## Control: lmerControl(optimizer = "bobyqa")
##
## REML criterion at convergence: 52555.5
##
## Scaled residuals:
## Min 1Q Median 3Q Max
## -2.4262 -0.6742 0.1455 0.7343 2.1944
##
## Random effects:
## Groups Name Variance Std.Dev. Corr
## country (Intercept) 0.7454 0.8634
## political_orientation 0.0329 0.1814 -0.84
## Residual 3.3402 1.8276
## Number of obs: 12949, groups: country, 144
##
## Fixed effects:
## Estimate Std. Error df t value Pr(>|t|)
## (Intercept) 3.40380 0.12195 69.80926 27.910 <2e-16 ***
## political_orientation -0.08334 0.03177 47.56677 -2.624 0.0117 *
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
##
## Correlation of Fixed Effects:
## (Intr)
## pltcl_rnttn -0.838## Computing profile confidence intervals ...## 0.15 % 99.85 %
## .sig01 0.5752608 1.24915249
## .sig02 -0.9536264 -0.49107070
## .sig03 0.1024774 0.29138678
## .sigma 1.7942564 1.86205701
## (Intercept) 3.0209153 3.76952829
## political_orientation -0.1784924 0.02029572Standardized Coefficients
## # A tibble: 2 × 5
## Parameter Std_Coefficient CI CI_low CI_high
## <chr> <dbl> <dbl> <dbl> <dbl>
## 1 (Intercept) -0.0173 0.997 -0.126 0.0912
## 2 political_orientation -0.0593 0.997 -0.126 0.00779H1a(2) Quadratic Effect
Here, we are examining the quadratic effect of right-wing political orientation on preferred political similarity in a partner using the Two Lines Approach (Simonsohn, 2018). We are using the Robin Hood Algorithm in order to set the breaking point. Then, we are calculating two multilevel regressions on either side of the breaking point. Outcome: Preference ratings for partner’s similar political beliefs and values. Predictors: Political Orientation & Age. Random intercept and random slope for country.
Algorithm: Find breaking point
See 11_twolines_analyses_multilevel
Regression 1 (x <= breaking_point)
model_pref_politicalsim_1 = lmer(pref_politicalsim ~ political_orientation +
(1+political_orientation|country),
data = data_included_documented %>%
dplyr::filter(political_orientation <= 3), control =lmerControl(optimizer = "bobyqa"))
summary(model_pref_politicalsim_1)## Linear mixed model fit by REML. t-tests use Satterthwaite's method [
## lmerModLmerTest]
## Formula:
## pref_politicalsim ~ political_orientation + (1 + political_orientation |
## country)
## Data: data_included_documented %>% dplyr::filter(political_orientation <=
## 3)
## Control: lmerControl(optimizer = "bobyqa")
##
## REML criterion at convergence: 42671.1
##
## Scaled residuals:
## Min 1Q Median 3Q Max
## -2.5773 -0.7032 0.1458 0.7426 2.2334
##
## Random effects:
## Groups Name Variance Std.Dev. Corr
## country (Intercept) 0.65527 0.8095
## political_orientation 0.05036 0.2244 -0.75
## Residual 3.18024 1.7833
## Number of obs: 10636, groups: country, 138
##
## Fixed effects:
## Estimate Std. Error df t value Pr(>|t|)
## (Intercept) 3.82477 0.12460 45.48408 30.70 < 2e-16 ***
## political_orientation -0.31340 0.04212 30.17206 -7.44 2.63e-08 ***
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
##
## Correlation of Fixed Effects:
## (Intr)
## pltcl_rnttn -0.815## Computing profile confidence intervals ...## 0.15 % 99.85 %
## .sig01 0.48282467 1.2567244
## .sig02 -0.92918790 -0.1896777
## .sig03 0.09638487 0.4008507
## .sigma 1.74741488 1.8206102
## (Intercept) 3.39772953 4.2064773
## political_orientation -0.44132932 -0.1574817## # A tibble: 2 × 5
## Parameter Std_Coefficient CI CI_low CI_high
## <chr> <dbl> <dbl> <dbl> <dbl>
## 1 (Intercept) 0 0.997 0 0
## 2 political_orientation -0.170 0.997 -0.237 -0.102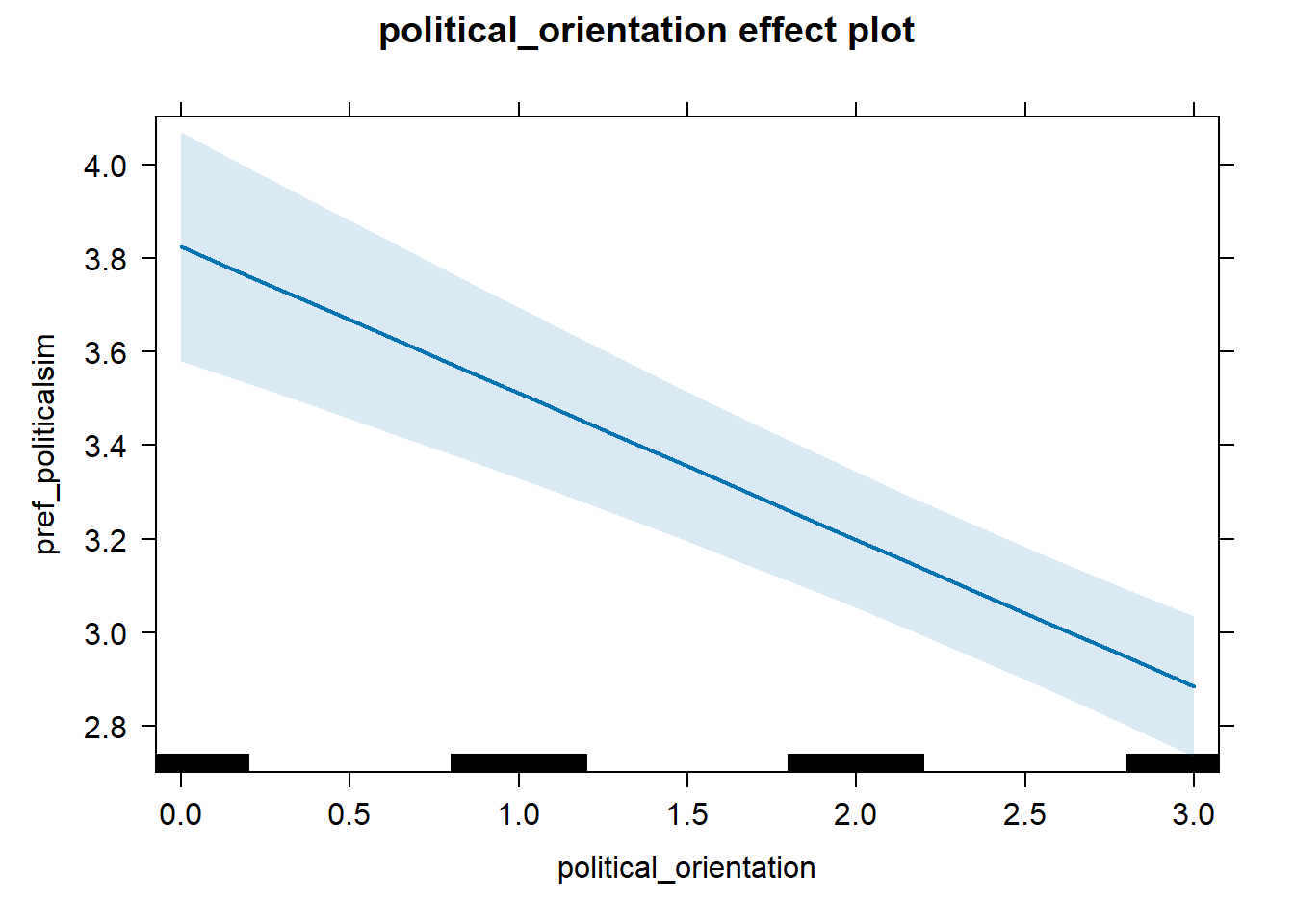
Regression 2 (x >= breaking_point)
model_pref_politicalsim_2 = lmer(pref_politicalsim ~ political_orientation +
(1+political_orientation|country),
data = data_included_documented %>%
dplyr::filter(political_orientation >= 3), control =lmerControl(optimizer = "bobyqa"))
summary(model_pref_politicalsim_2)## Linear mixed model fit by REML. t-tests use Satterthwaite's method [
## lmerModLmerTest]
## Formula:
## pref_politicalsim ~ political_orientation + (1 + political_orientation |
## country)
## Data: data_included_documented %>% dplyr::filter(political_orientation >=
## 3)
## Control: lmerControl(optimizer = "bobyqa")
##
## REML criterion at convergence: 29821.8
##
## Scaled residuals:
## Min 1Q Median 3Q Max
## -2.8134 -0.7937 0.1168 0.6892 2.1899
##
## Random effects:
## Groups Name Variance Std.Dev. Corr
## country (Intercept) 0.7483 0.8650
## political_orientation 0.0314 0.1772 -0.81
## Residual 3.3136 1.8203
## Number of obs: 7357, groups: country, 129
##
## Fixed effects:
## Estimate Std. Error df t value Pr(>|t|)
## (Intercept) 1.62534 0.17798 25.66396 9.132 1.53e-09 ***
## political_orientation 0.40821 0.04567 22.90045 8.939 6.31e-09 ***
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
##
## Correlation of Fixed Effects:
## (Intr)
## pltcl_rnttn -0.911## Computing profile confidence intervals ...## 0.15 % 99.85 %
## .sig01 0.32893449 1.6393578
## .sig02 -0.97195114 0.4328139
## .sig03 0.04045946 0.3781595
## .sigma 1.77636620 1.8662207
## (Intercept) 1.08787002 2.3016652
## political_orientation 0.23113547 0.5549943## # A tibble: 2 × 5
## Parameter Std_Coefficient CI CI_low CI_high
## <chr> <dbl> <dbl> <dbl> <dbl>
## 1 (Intercept) 0 0.997 0 0
## 2 political_orientation 0.175 0.997 0.117 0.233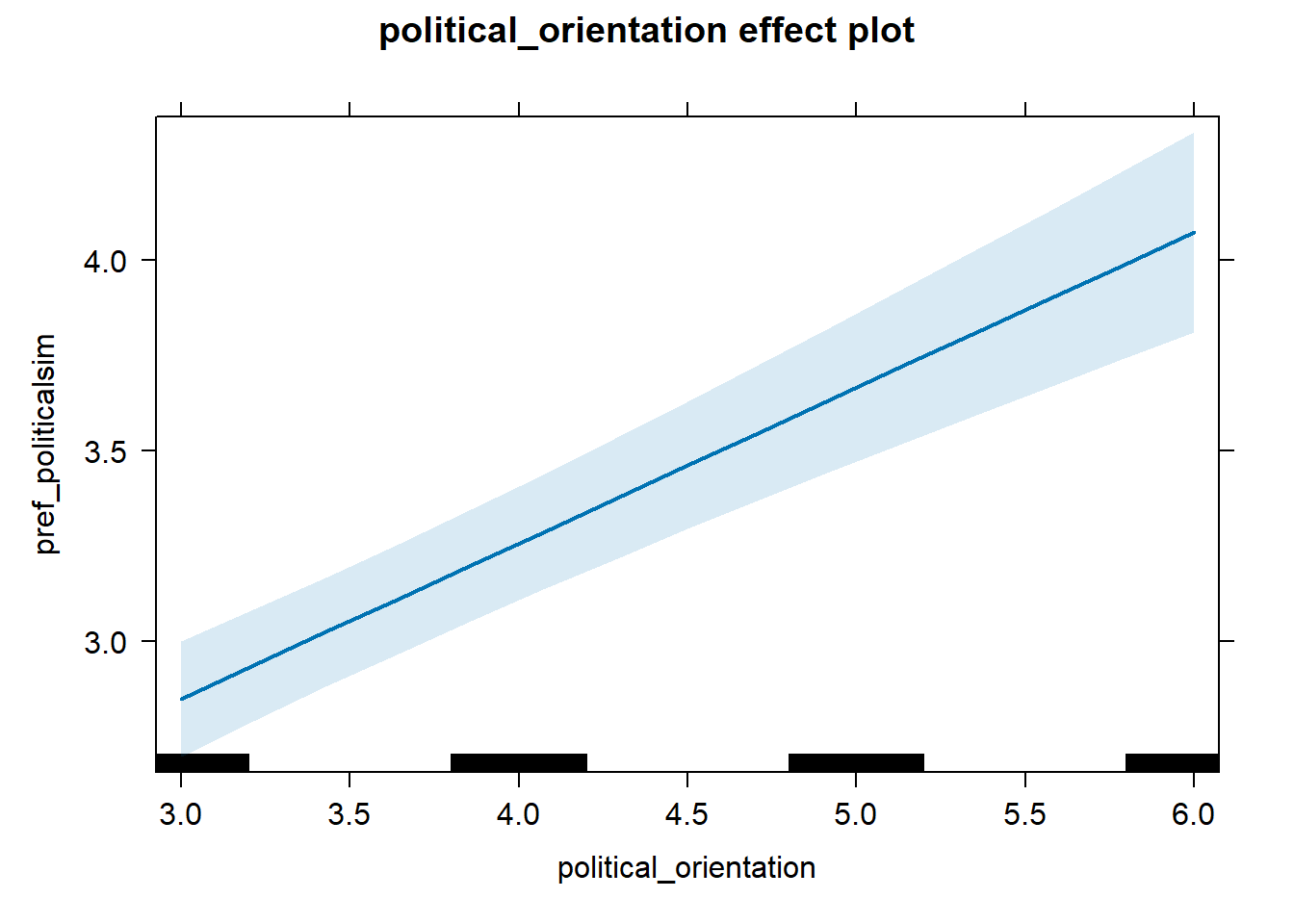
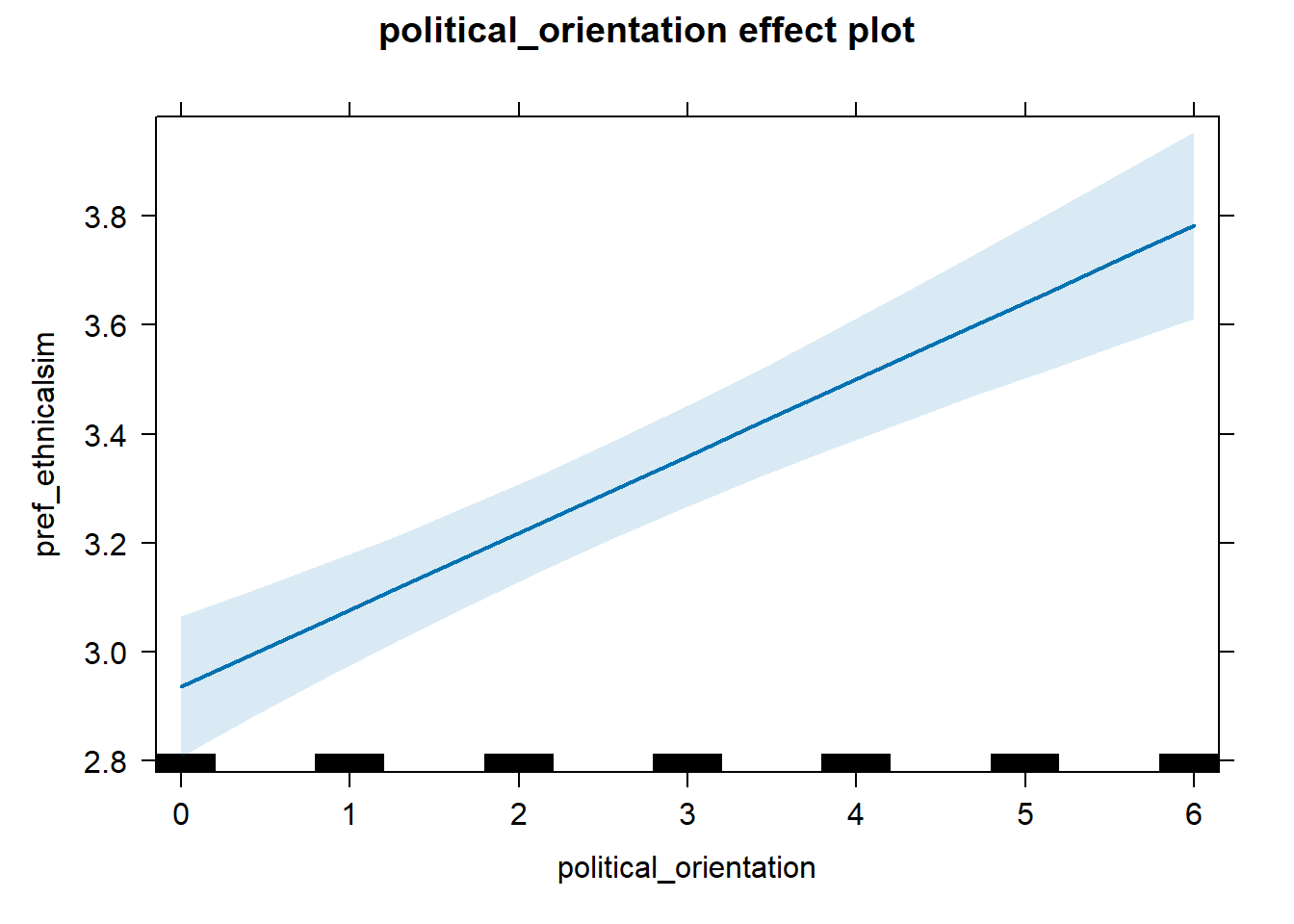
H1b Preference for Similarity in Ethnicity/Race
H1b There is a positive linear link between right-wing political orientation and women’s preferences for partner’s similar ethnicity/race. Outcome: Preference ratings for partner’s similar ethnicity/race. Predictor: Political Orientation. Random intercept and random slope for country.
Models
Summary
## Linear mixed model fit by REML. t-tests use Satterthwaite's method [
## lmerModLmerTest]
## Formula:
## pref_ethnicalsim ~ political_orientation + (1 + political_orientation |
## country)
## Data: data_included_documented
## Control: lmerControl(optimizer = "bobyqa")
##
## REML criterion at convergence: 30431.4
##
## Scaled residuals:
## Min 1Q Median 3Q Max
## -3.2873 -0.4577 -0.0320 0.5952 2.4045
##
## Random effects:
## Groups Name Variance Std.Dev. Corr
## country (Intercept) 0.083581 0.28910
## political_orientation 0.006438 0.08024 -0.40
## Residual 1.955963 1.39856
## Number of obs: 8643, groups: country, 127
##
## Fixed effects:
## Estimate Std. Error df t value Pr(>|t|)
## (Intercept) 2.93636 0.06571 24.05651 44.689 < 2e-16 ***
## political_orientation 0.14098 0.02047 25.38692 6.889 2.95e-07 ***
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
##
## Correlation of Fixed Effects:
## (Intr)
## pltcl_rnttn -0.728## Computing profile confidence intervals ...## 0.15 % 99.85 %
## .sig01 0.11069563 0.5700297
## .sig02 -0.87971422 0.7945738
## .sig03 0.03014798 0.1653320
## .sigma 1.36741251 1.4309525
## (Intercept) 2.73858107 3.1632222
## political_orientation 0.07038154 0.2025605Standardized Coefficients
## # A tibble: 2 × 5
## Parameter Std_Coefficient CI CI_low CI_high
## <chr> <dbl> <dbl> <dbl> <dbl>
## 1 (Intercept) 0 0.997 0 0
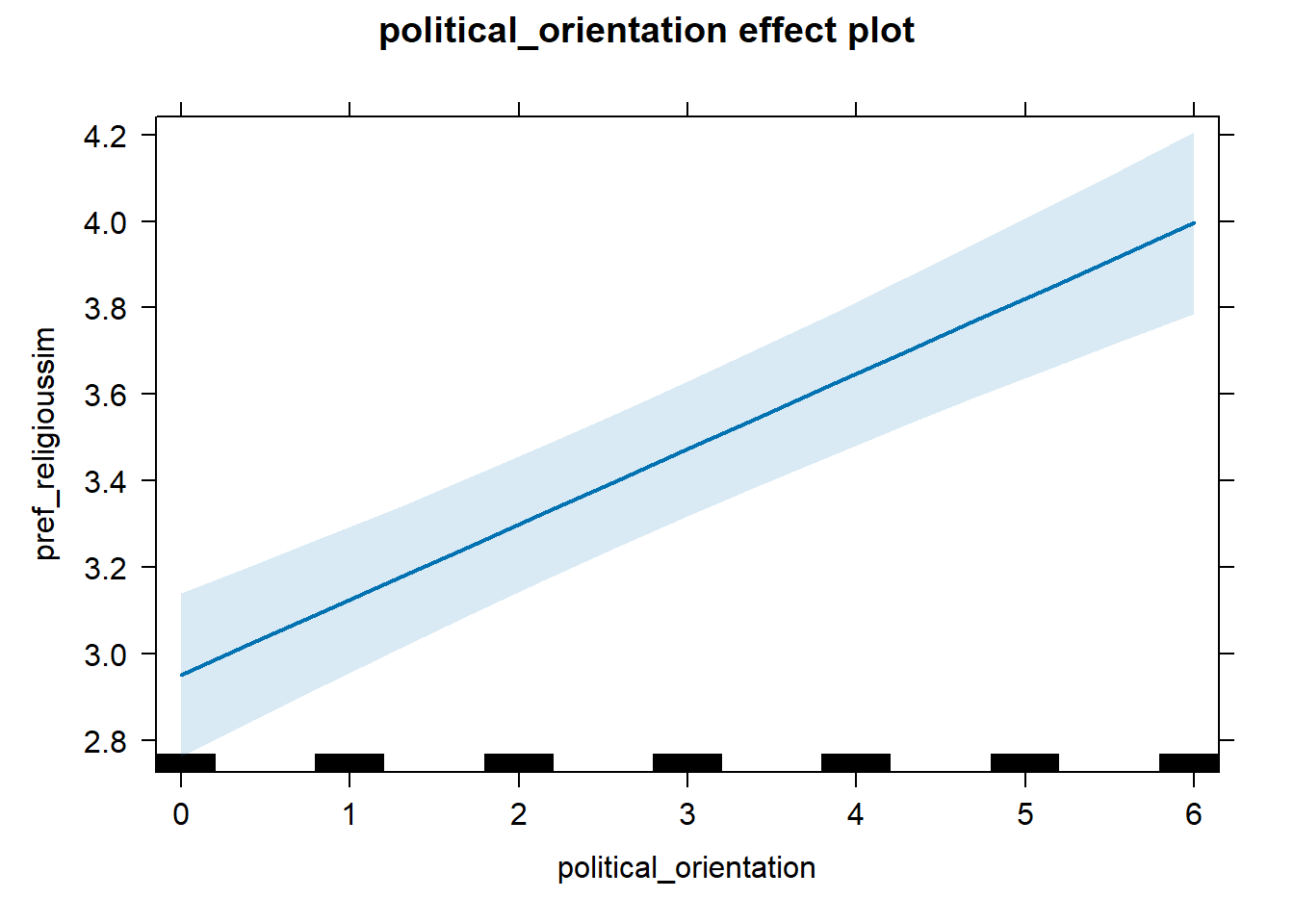
## 2 political_orientation 0.128 0.997 0.0731 0.184H1c Preference for Similarity in Religion
H1c There is a positive linear link between right-wing political
orientation and women’s preferences for partner’s similar religious
beliefs.
Outcome: Preference ratings for partner’s similar religious beliefs.
Predictor: Political Orientation. Random intercept and random slope for
country.
Models
Summary
## Linear mixed model fit by REML. t-tests use Satterthwaite's method [
## lmerModLmerTest]
## Formula:
## pref_religioussim ~ political_orientation + (1 + political_orientation |
## country)
## Data: data_included_documented
## Control: lmerControl(optimizer = "bobyqa")
##
## REML criterion at convergence: 56523.3
##
## Scaled residuals:
## Min 1Q Median 3Q Max
## -2.1900 -1.0116 0.0650 0.8719 1.9319
##
## Random effects:
## Groups Name Variance Std.Dev. Corr
## country (Intercept) 0.389003 0.62370
## political_orientation 0.004839 0.06956 -0.29
## Residual 4.571227 2.13804
## Number of obs: 12937, groups: country, 144
##
## Fixed effects:
## Estimate Std. Error df t value Pr(>|t|)
## (Intercept) 2.95120 0.09640 31.33217 30.614 < 2e-16 ***
## political_orientation 0.17408 0.02132 23.38888 8.166 2.66e-08 ***
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
##
## Correlation of Fixed Effects:
## (Intr)
## pltcl_rnttn -0.575## Computing profile confidence intervals ...## 0.15 % 99.85 %
## .sig01 0.309523959 1.0217923
## .sig02 -0.907182380 0.9735600
## .sig03 0.004622921 0.1587600
## .sigma 2.099015572 2.1783470
## (Intercept) 2.637243786 3.3410490
## political_orientation 0.073780282 0.2632057Standardized Coefficients
## # A tibble: 2 × 5
## Parameter Std_Coefficient CI CI_low CI_high
## <chr> <dbl> <dbl> <dbl> <dbl>
## 1 (Intercept) 0 0.997 0 0
## 2 political_orientation 0.108 0.997 0.0686 0.147Ideal Partner Preferences
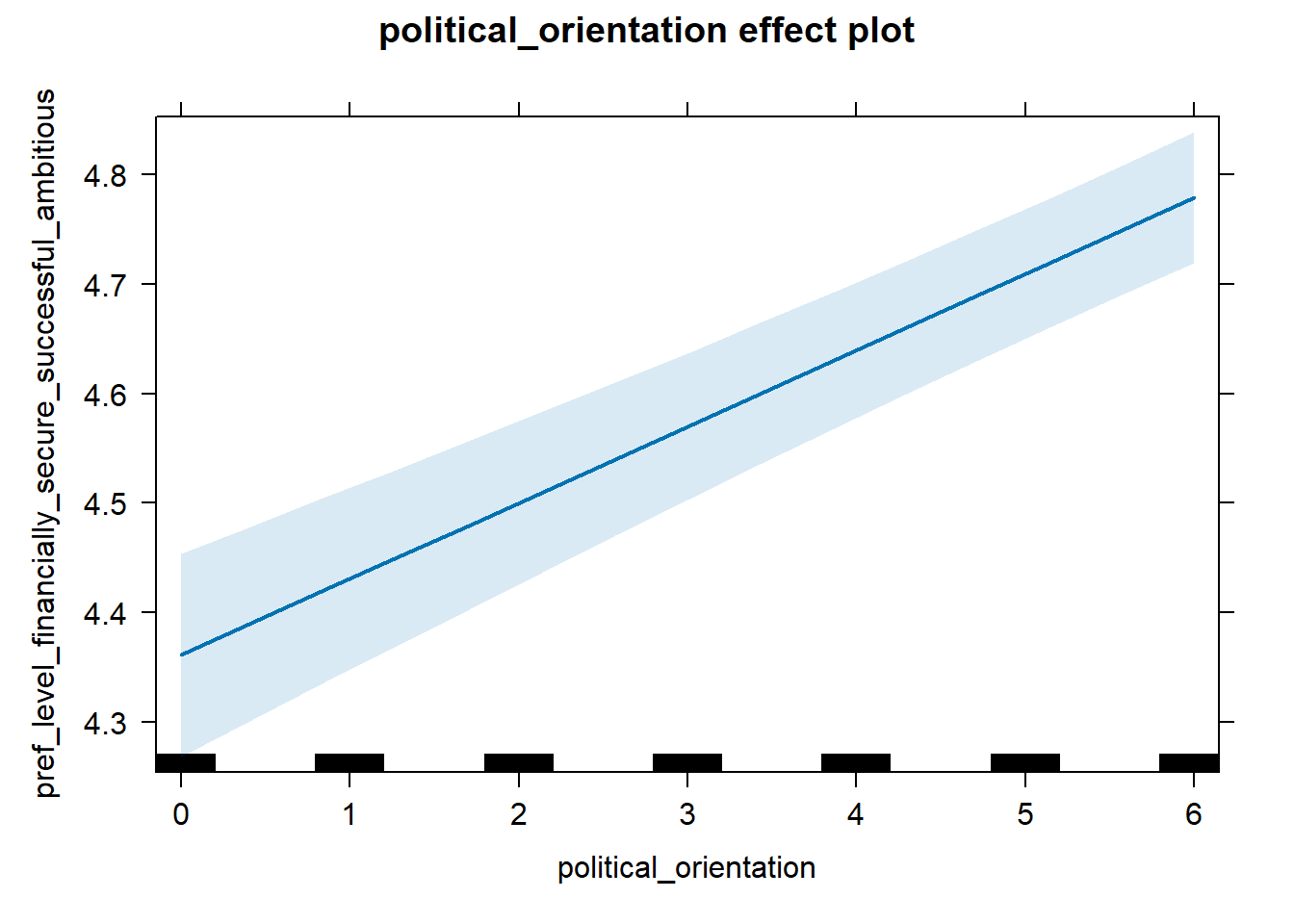
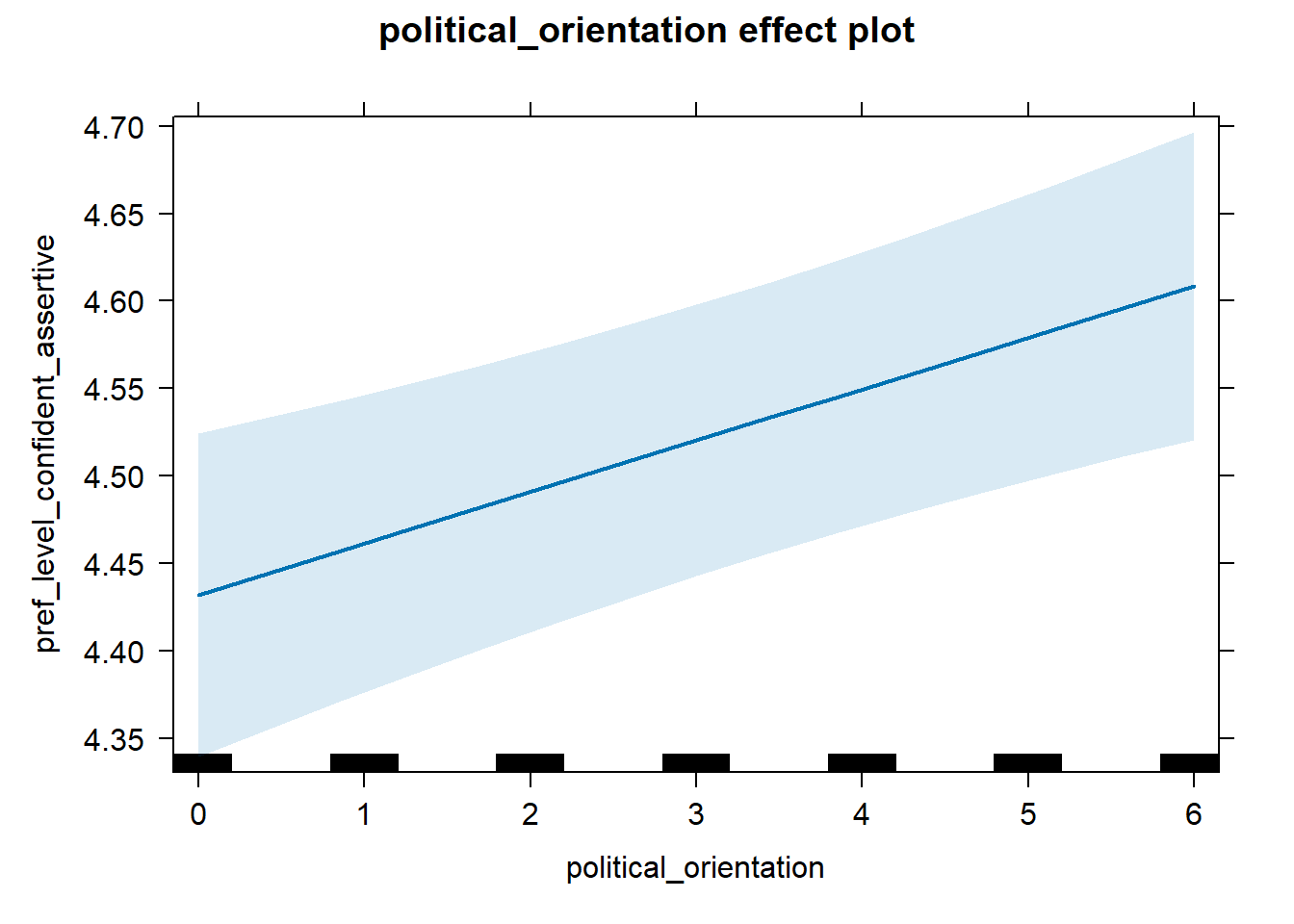
H2a Preference for the Level of Financial Security- Successfulness
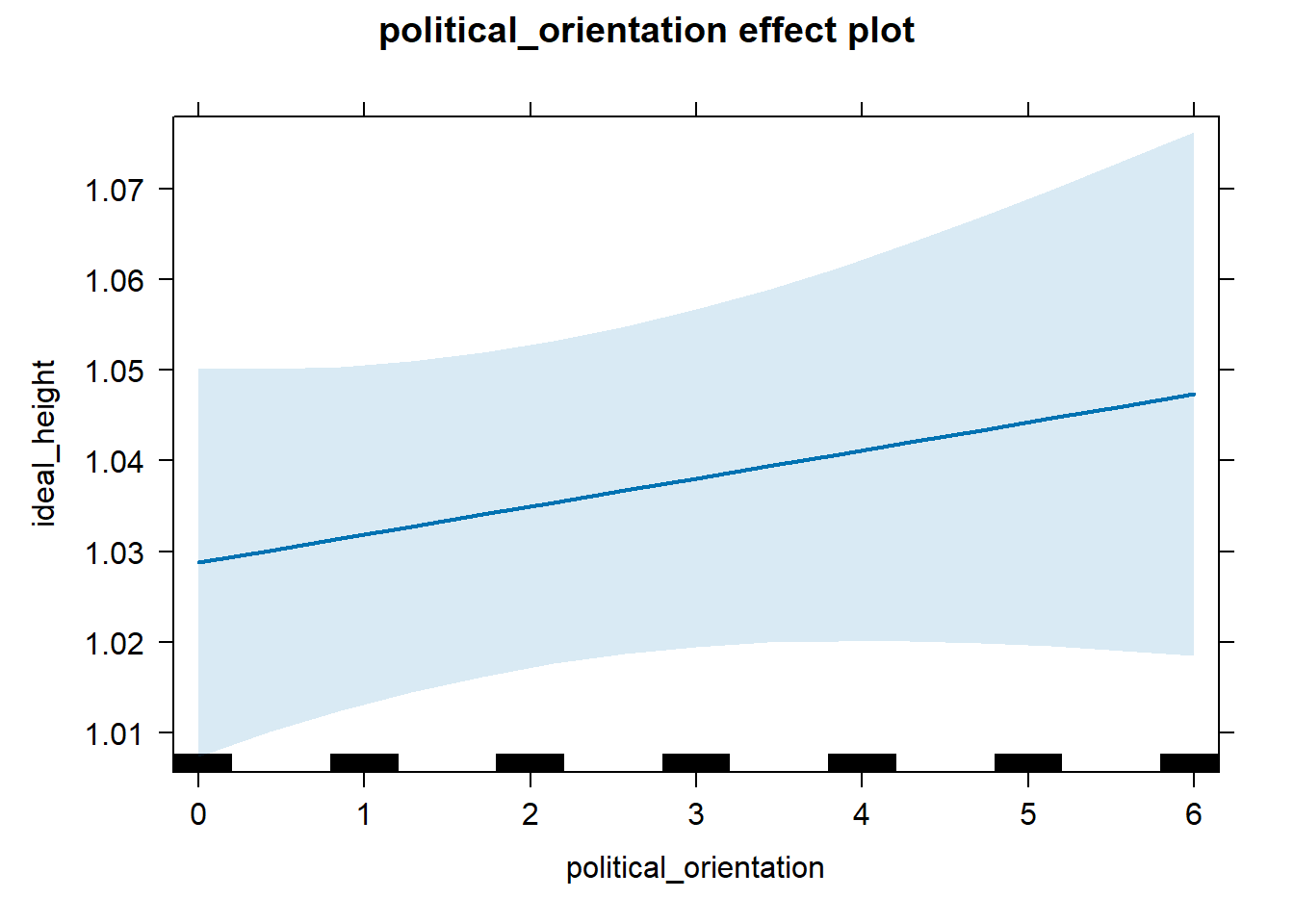
H2a There is a positive linear link between right-wing political orientation and women’s preferences for the level of financial security and successfulness. Outcome: Level ratings for partner’s financial security-successfulness. Predictor: Political Orientation. Random intercept and random slope for country.
Models
model_pref_level_financially_secure_successful_ambitious <-
lmer(
pref_level_financially_secure_successful_ambitious ~ political_orientation + (1 +
political_orientation |
country),
data = data_included_documented,
control = lmerControl(optimizer = "bobyqa")
)## boundary (singular) fit: see help('isSingular')Summary
## Linear mixed model fit by REML. t-tests use Satterthwaite's method [
## lmerModLmerTest]
## Formula:
## pref_level_financially_secure_successful_ambitious ~ political_orientation +
## (1 + political_orientation | country)
## Data: data_included_documented
## Control: lmerControl(optimizer = "bobyqa")
##
## REML criterion at convergence: 31715.7
##
## Scaled residuals:
## Min 1Q Median 3Q Max
## -4.8198 -0.7049 0.0118 0.7074 2.5653
##
## Random effects:
## Groups Name Variance Std.Dev. Corr
## country (Intercept) 0.1340500 0.36613
## political_orientation 0.0009219 0.03036 -1.00
## Residual 0.7234663 0.85057
## Number of obs: 12551, groups: country, 143
##
## Fixed effects:
## Estimate Std. Error df t value Pr(>|t|)
## (Intercept) 4.360909 0.047409 75.507204 91.99 <2e-16 ***
## political_orientation 0.069690 0.006831 99.425822 10.20 <2e-16 ***
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
##
## Correlation of Fixed Effects:
## (Intr)
## pltcl_rnttn -0.771
## optimizer (bobyqa) convergence code: 0 (OK)
## boundary (singular) fit: see help('isSingular')## Computing profile confidence intervals ...## Warning in zeta(shiftpar, start = opt[seqpar1][-w]): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in FUN(X[[i]], ...): non-monotonic profile for .sig01## Warning in FUN(X[[i]], ...): non-monotonic profile for .sig02## Warning in zeta(shiftpar, start = opt[seqpar1][-w]): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in FUN(X[[i]], ...): non-monotonic profile for .sig03## Warning in zeta(shiftpar, start = opt[seqpar1][-w]): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in zetafun(np, ns): NAs detected in profiling## Warning in FUN(X[[i]], ...): non-monotonic profile for .sigma## Warning in min(diff(obj1[, 2]) < (-non.mono.tol), na.rm = TRUE): kein
## nicht-fehlendes Argument für min; gebe Inf zurück## Warning in confint.thpr(pp, level = level, zeta = zeta): non-monotonic profile
## for .sig01## Warning in confint.thpr(pp, level = level, zeta = zeta): bad spline fit for
## .sig02: falling back to linear interpolation## Warning in min(diff(obj1[, 2]) < (-non.mono.tol), na.rm = TRUE): kein
## nicht-fehlendes Argument für min; gebe Inf zurück## Warning in confint.thpr(pp, level = level, zeta = zeta): non-monotonic profile
## for .sig03## Warning in min(diff(obj1[, 2]) < (-non.mono.tol), na.rm = TRUE): kein
## nicht-fehlendes Argument für min; gebe Inf zurück## Warning in confint.thpr(pp, level = level, zeta = zeta): non-monotonic profile
## for .sigma## 0.15 % 99.85 %
## .sig01 NA NA
## .sig02 -1.00639456 1.00000000
## .sig03 NA NA
## .sigma NA NA
## (Intercept) 4.21955176 4.51794888
## political_orientation 0.04395714 0.09167436Standardized Coefficients
standardize_parameters(model_pref_level_financially_secure_successful_ambitious, method = "basic", ci = 0.997)## # A tibble: 2 × 5
## Parameter Std_Coefficient CI CI_low CI_high
## <chr> <dbl> <dbl> <dbl> <dbl>
## 1 (Intercept) 0 0.997 0 0
## 2 political_orientation 0.105 0.997 0.0747 0.136H2b Preference for the Level of Confidence-Assertiveness
H2b There is a positive linear link between right-wing political
orientation and women’s preferences for the level of confidence and
assertiveness.
Outcome: Level ratings for partner’s confidence-assertiveness.
Predictor: Political Orientation. Random intercept and random slope for
country.
Models
Summary
## Linear mixed model fit by REML. t-tests use Satterthwaite's method [
## lmerModLmerTest]
## Formula: pref_level_confident_assertive ~ political_orientation + (1 +
## political_orientation | country)
## Data: data_included_documented
## Control: lmerControl(optimizer = "bobyqa")
##
## REML criterion at convergence: 28928.4
##
## Scaled residuals:
## Min 1Q Median 3Q Max
## -5.4786 -0.6798 -0.0354 0.6526 3.0002
##
## Random effects:
## Groups Name Variance Std.Dev. Corr
## country (Intercept) 0.1518032 0.38962
## political_orientation 0.0007535 0.02745 -0.52
## Residual 0.5614583 0.74931
## Number of obs: 12697, groups: country, 144
##
## Fixed effects:
## Estimate Std. Error df t value Pr(>|t|)
## (Intercept) 4.432e+00 4.713e-02 1.022e+02 94.044 < 2e-16 ***
## political_orientation 2.946e-02 7.797e-03 2.185e+01 3.778 0.00104 **
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
##
## Correlation of Fixed Effects:
## (Intr)
## pltcl_rnttn -0.544## Computing profile confidence intervals ...## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): unexpected decrease in
## profile: using minstep## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in FUN(X[[i]], ...): non-monotonic profile for .sig02## Warning in confint.thpr(pp, level = level, zeta = zeta): bad spline fit for
## .sig02: falling back to linear interpolation## Warning in regularize.values(x, y, ties, missing(ties), na.rm = na.rm):
## Reduktion auf einmalige 'x' Werte## 0.15 % 99.85 %
## .sig01 0.288700584 0.52594291
## .sig02 -0.967689483 0.23965114
## .sig03 0.003602748 0.05946370
## .sigma 0.735494367 0.76355617
## (Intercept) 4.291019618 4.57603976
## political_orientation 0.003430270 0.05344236Standardized Coefficients
## # A tibble: 2 × 5
## Parameter Std_Coefficient CI CI_low CI_high
## <chr> <dbl> <dbl> <dbl> <dbl>
## 1 (Intercept) 0 0.997 0 0
## 2 political_orientation 0.0476 0.997 0.0102 0.0850H2c Preference for the Level of Education-Intelligence
H2c There is no link between right-wing political orientation and
women’s preferences for the level of education and intelligence.
Outcome: Level ratings for partner’s education-intelligence. Predictor:
Political Orientation. Random intercept and random slope for
country.
Models
Summary
## Linear mixed model fit by REML. t-tests use Satterthwaite's method [
## lmerModLmerTest]
## Formula: pref_level_intelligence_educated ~ political_orientation + (1 +
## political_orientation | country)
## Data: data_included_documented
## Control: lmerControl(optimizer = "bobyqa")
##
## REML criterion at convergence: 30133.6
##
## Scaled residuals:
## Min 1Q Median 3Q Max
## -5.2463 -0.6840 0.0410 0.6903 2.1702
##
## Random effects:
## Groups Name Variance Std.Dev. Corr
## country (Intercept) 0.0627259 0.25045
## political_orientation 0.0009513 0.03084 -0.22
## Residual 0.6171382 0.78558
## Number of obs: 12723, groups: country, 141
##
## Fixed effects:
## Estimate Std. Error df t value Pr(>|t|)
## (Intercept) 4.80150 0.03776 71.33658 127.162 <2e-16 ***
## political_orientation 0.02120 0.00859 28.58180 2.468 0.0198 *
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
##
## Correlation of Fixed Effects:
## (Intr)
## pltcl_rnttn -0.556## Computing profile confidence intervals ...## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): Last two rows have
## identical or NA .zeta values: using minstep## Warning in FUN(X[[i]], ...): non-monotonic profile for .sig02## Warning in confint.thpr(pp, level = level, zeta = zeta): bad spline fit for
## .sig02: falling back to linear interpolation## Warning in regularize.values(x, y, ties, missing(ties), na.rm = na.rm):
## Reduktion auf einmalige 'x' Werte## 0.15 % 99.85 %
## .sig01 0.167528212 0.36538714
## .sig02 -0.796184160 0.59712860
## .sig03 0.009011258 0.06387362
## .sigma 0.771123896 0.80048216
## (Intercept) 4.687218265 4.92338841
## political_orientation -0.006088975 0.05130632Standardized Coefficients
## # A tibble: 2 × 5
## Parameter Std_Coefficient CI CI_low CI_high
## <chr> <dbl> <dbl> <dbl> <dbl>
## 1 (Intercept) 0 0.997 0 0
## 2 political_orientation 0.0351 0.997 -0.00712 0.0774Plot
lmer(pref_level_intelligence_educated ~ political_orientation + (1+political_orientation|country),
data = data_included_documented) %>%
allEffects() %>%
plot()## Warning in checkConv(attr(opt, "derivs"), opt$par, ctrl = control$checkConv, :
## Model failed to converge with max|grad| = 0.00200785 (tol = 0.002, component 1)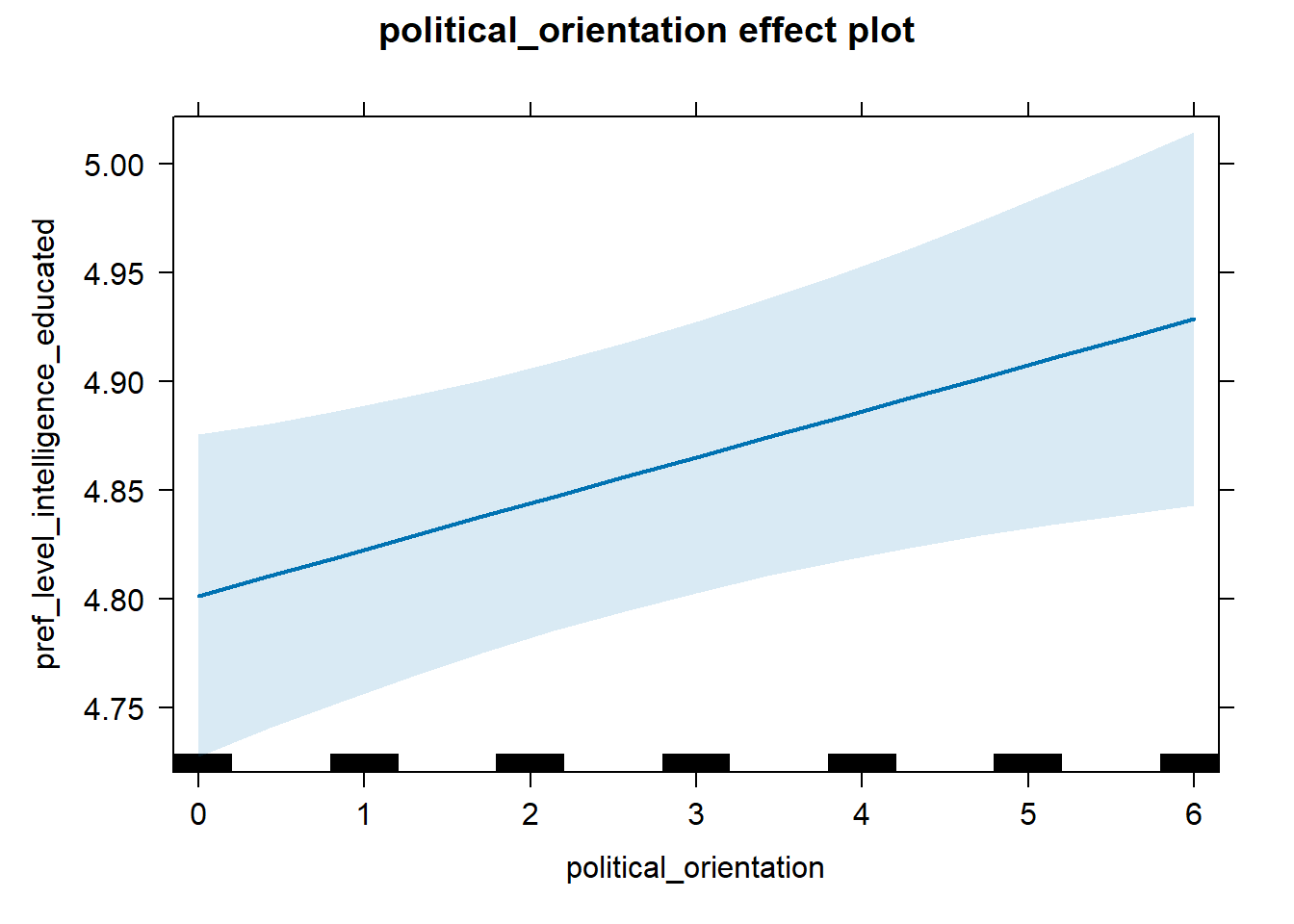
H2d Preference for the Level of Kindness-Supportiveness
H2d There is no link between right-wing political orientation and
women’s preferences for the level of kindness and supportiveness.
Outcome: Level ratings for partner’s kindness-supportiveness. Predictor:
Political Orientation. Random intercept and random slope for
country.
Models
Summary
## Linear mixed model fit by REML. t-tests use Satterthwaite's method [
## lmerModLmerTest]
## Formula:
## pref_level_kind_supportive ~ political_orientation + (1 + political_orientation |
## country)
## Data: data_included_documented
## Control: lmerControl(optimizer = "bobyqa")
##
## REML criterion at convergence: 26550.1
##
## Scaled residuals:
## Min 1Q Median 3Q Max
## -4.7907 -0.6652 0.0687 0.8000 1.7293
##
## Random effects:
## Groups Name Variance Std.Dev. Corr
## country (Intercept) 1.657e-02 0.128718
## political_orientation 5.642e-05 0.007511 0.56
## Residual 4.674e-01 0.683662
## Number of obs: 12729, groups: country, 143
##
## Fixed effects:
## Estimate Std. Error df t value Pr(>|t|)
## (Intercept) 5.163681 0.022796 44.855531 226.521 <2e-16 ***
## political_orientation 0.011053 0.004833 14.568403 2.287 0.0376 *
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
##
## Correlation of Fixed Effects:
## (Intr)
## pltcl_rnttn -0.426## Computing profile confidence intervals ...## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): unexpected decrease in
## profile: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): unexpected decrease in
## profile: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): unexpected decrease in
## profile: using minstep## Warning in FUN(X[[i]], ...): non-monotonic profile for .sig01## Warning in FUN(X[[i]], ...): non-monotonic profile for .sig02## Warning in confint.thpr(pp, level = level, zeta = zeta): bad spline fit for
## .sig01: falling back to linear interpolation## Warning in confint.thpr(pp, level = level, zeta = zeta): bad spline fit for
## .sig02: falling back to linear interpolation## 0.15 % 99.85 %
## .sig01 0.085731705 0.20670347
## .sig02 -1.000000000 1.00000000
## .sig03 0.000000000 0.03294017
## .sigma 0.671097562 0.69661278
## (Intercept) 5.095444447 5.23771219
## political_orientation -0.005103504 0.02736140Standardized Coefficients
## # A tibble: 2 × 5
## Parameter Std_Coefficient CI CI_low CI_high
## <chr> <dbl> <dbl> <dbl> <dbl>
## 1 (Intercept) 0 0.997 0 0
## 2 political_orientation 0.0215 0.997 -0.00640 0.0494Plot
lmer(pref_level_kind_supportive ~ political_orientation + (1+political_orientation|country),
data = data_included_documented) %>%
allEffects() %>%
plot()## Warning in checkConv(attr(opt, "derivs"), opt$par, ctrl = control$checkConv, :
## Model failed to converge with max|grad| = 0.00451923 (tol = 0.002, component 1)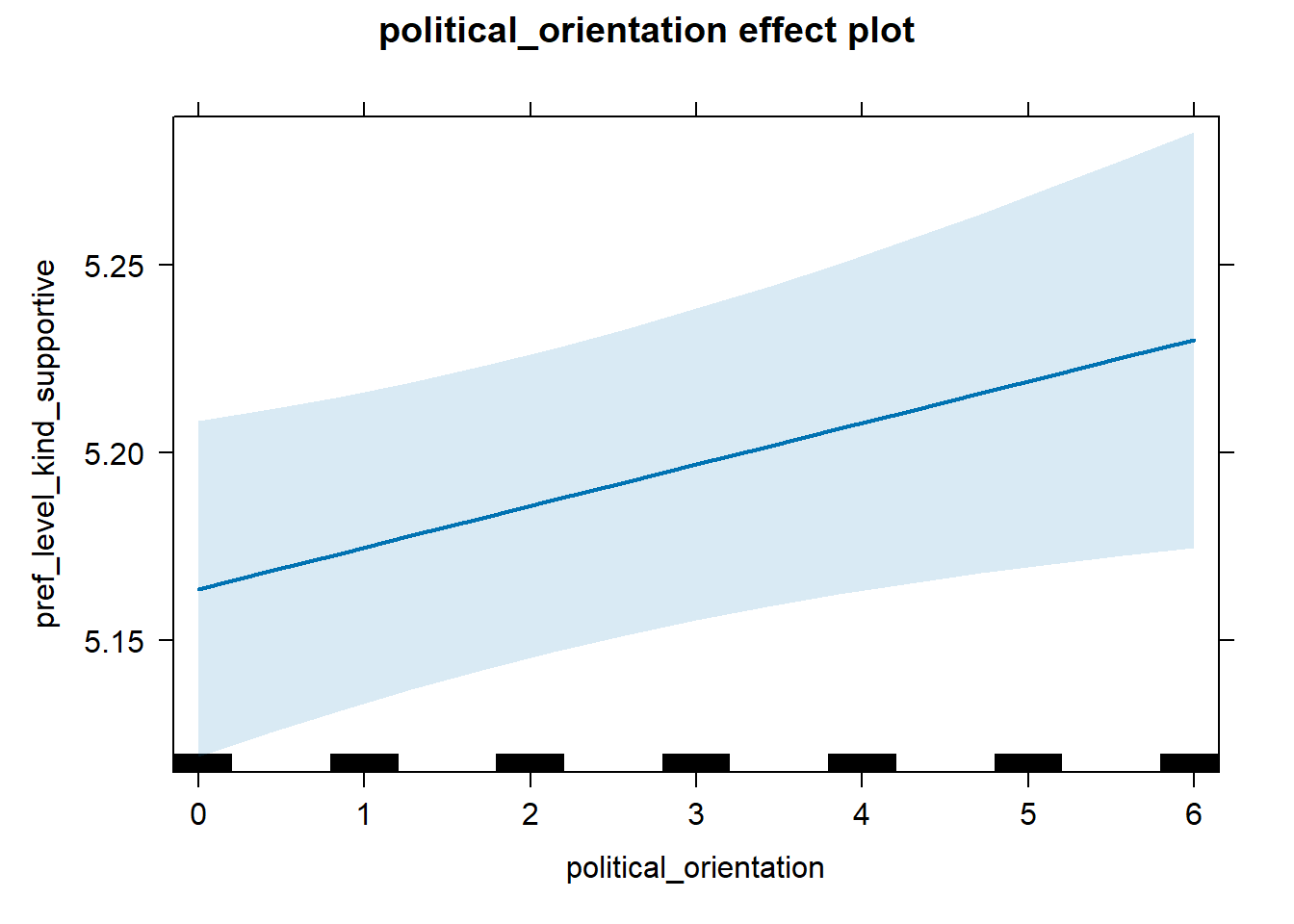
H2e Preference for the Level of Attractiveness
H2e There is no link between right-wing political orientation and
women’s preferences for the level of attractiveness.
Outcome: Level ratings for partner’s attractiveness. Predictor:
Political Orientation. Random intercept and random slope for
country.
Models
Summary
## Linear mixed model fit by REML. t-tests use Satterthwaite's method [
## lmerModLmerTest]
## Formula:
## pref_level_attractiveness ~ political_orientation + (1 + political_orientation |
## country)
## Data: data_included_documented
## Control: lmerControl(optimizer = "bobyqa")
##
## REML criterion at convergence: 32918.9
##
## Scaled residuals:
## Min 1Q Median 3Q Max
## -4.6503 -0.6477 -0.0244 0.6476 2.5185
##
## Random effects:
## Groups Name Variance Std.Dev. Corr
## country (Intercept) 0.01093 0.10452
## political_orientation 0.00027 0.01643 0.28
## Residual 0.80567 0.89759
## Number of obs: 12526, groups: country, 142
##
## Fixed effects:
## Estimate Std. Error df t value Pr(>|t|)
## (Intercept) 3.927775 0.025446 33.185806 154.356 < 2e-16 ***
## political_orientation 0.039238 0.007106 12.674681 5.522 0.000108 ***
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
##
## Correlation of Fixed Effects:
## (Intr)
## pltcl_rnttn -0.559## Computing profile confidence intervals ...## 0.15 % 99.85 %
## .sig01 0.06770729 0.18832187
## .sig02 -0.75850240 0.91589442
## .sig03 0.00000000 0.05867958
## .sigma 0.88097657 0.91472753
## (Intercept) 3.84943833 4.00914319
## political_orientation 0.01356438 0.06154413Standardized Coefficients
## # A tibble: 2 × 5
## Parameter Std_Coefficient CI CI_low CI_high
## <chr> <dbl> <dbl> <dbl> <dbl>
## 1 (Intercept) 0 0.997 0 0
## 2 political_orientation 0.0583 0.997 0.0270 0.0897Plot
lmer(pref_level_attractiveness ~ political_orientation + (1+political_orientation|country),
data = data_included_documented) %>%
allEffects() %>%
plot()## Warning in checkConv(attr(opt, "derivs"), opt$par, ctrl = control$checkConv, :
## Model failed to converge with max|grad| = 0.002634 (tol = 0.002, component 1)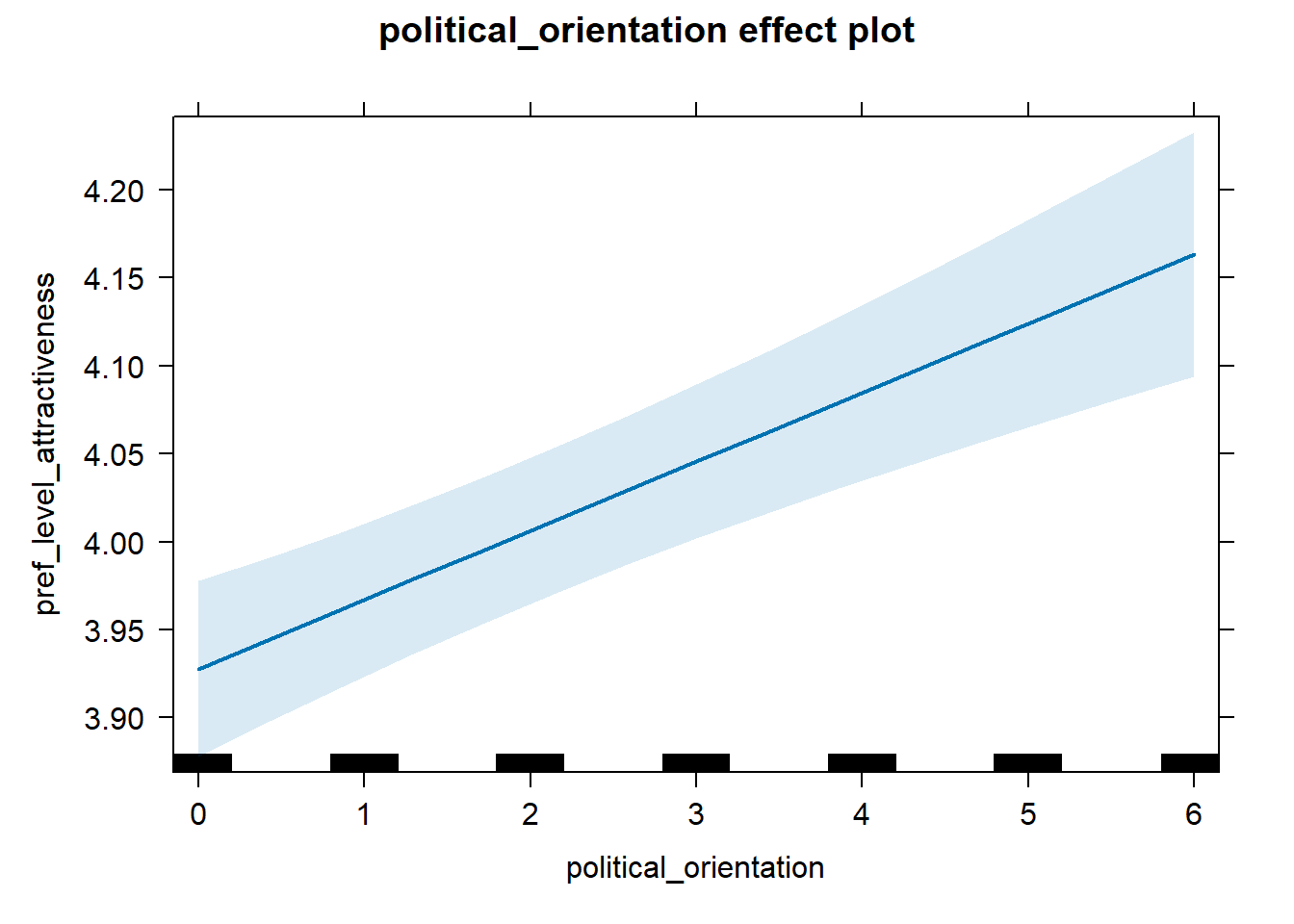
Ideal Age and Height
H3a(1) Importance Ratings for Partner’s Age
H3a(1) There is a positive linear link between right-wing political orientation and women’s importance ratings for partner’s age. Outcome: Importance ratings for partner’s age. Predictor: Political Orientation. Random intercept and random slope for country.
Models
Summary
## Linear mixed model fit by REML. t-tests use Satterthwaite's method [
## lmerModLmerTest]
## Formula: imp_age ~ political_orientation + (1 + political_orientation |
## country)
## Data: data_included_documented
## Control: lmerControl(optimizer = "bobyqa")
##
## REML criterion at convergence: 46141.4
##
## Scaled residuals:
## Min 1Q Median 3Q Max
## -3.0525 -0.5057 0.1715 0.6392 2.1583
##
## Random effects:
## Groups Name Variance Std.Dev. Corr
## country (Intercept) 0.08563 0.2926
## political_orientation 0.01025 0.1012 -0.76
## Residual 1.95847 1.3995
## Number of obs: 13113, groups: country, 143
##
## Fixed effects:
## Estimate Std. Error df t value Pr(>|t|)
## (Intercept) 3.50977 0.05990 21.71636 58.591 < 2e-16 ***
## political_orientation 0.08199 0.02058 15.91106 3.985 0.00108 **
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
##
## Correlation of Fixed Effects:
## (Intr)
## pltcl_rnttn -0.832## Computing profile confidence intervals ...## 0.15 % 99.85 %
## .sig01 0.108667718 0.5631709
## .sig02 -0.960382731 0.7688915
## .sig03 0.009474955 0.2058263
## .sigma 1.374118745 1.4257678
## (Intercept) 3.320129690 3.7190380
## political_orientation 0.013398755 0.1510101Standardized Coefficients
## # A tibble: 2 × 5
## Parameter Std_Coefficient CI CI_low CI_high
## <chr> <dbl> <dbl> <dbl> <dbl>
## 1 (Intercept) 0 0.997 0 0
## 2 political_orientation 0.0787 0.997 0.0201 0.137H3a(2) Level Ratings for Partner’s Age
H3a(2) There is a positive linear link between right-wing political orientation and the relative age discrepancy between ideal partner’s age and women’s age. (discrepancy calculated as ideal partner’s age – women’s age) Outcome: Discrepancy between level ratings for ideal partner’s age and women’s age (ideal_age_rel) Predictor: Political Orientation. Random intercept and random slope for country.
Models
Summary
## Linear mixed model fit by REML. t-tests use Satterthwaite's method [
## lmerModLmerTest]
## Formula: ideal_age_rel ~ political_orientation + (1 + political_orientation |
## country)
## Data: data_included_documented
## Control: lmerControl(optimizer = "bobyqa")
##
## REML criterion at convergence: 61688.6
##
## Scaled residuals:
## Min 1Q Median 3Q Max
## -11.6404 -0.4618 -0.1175 0.4395 25.7025
##
## Random effects:
## Groups Name Variance Std.Dev. Corr
## country (Intercept) 0.8873 0.9420
## political_orientation 0.4705 0.6859 -0.94
## Residual 9.5961 3.0978
## Number of obs: 12057, groups: country, 140
##
## Fixed effects:
## Estimate Std. Error df t value Pr(>|t|)
## (Intercept) 2.39574 0.14881 14.23277 16.099 1.56e-10 ***
## political_orientation 0.15044 0.08541 35.89473 1.761 0.0867 .
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
##
## Correlation of Fixed Effects:
## (Intr)
## pltcl_rnttn -0.871## Computing profile confidence intervals ...## 0.15 % 99.85 %
## .sig01 0.2473102 1.931245124
## .sig02 -0.9925466 0.009045051
## .sig03 0.4001037 1.029585987
## .sigma 3.0391523 3.158354644
## (Intercept) 1.8588095 2.891544423
## political_orientation -0.1136682 0.420746805Standardized Coefficients
## # A tibble: 2 × 5
## Parameter Std_Coefficient CI CI_low CI_high
## <chr> <dbl> <dbl> <dbl> <dbl>
## 1 (Intercept) 0 0.997 0 0
## 2 political_orientation 0.0652 0.997 -0.0447 0.175H3b(1) Importance Ratings for Partner’s Height
H3b(1) There is a positive linear link between right-wing political orientation and women’s importance ratings for partner’s height. Outcome: Importance ratings for partner’s height. Predictor: Political Orientation. Random intercept and random slope for country.
Models
model_imp_height <- lmer(imp_height ~ political_orientation + (1+political_orientation|country), data = data_included_documented, control =lmerControl(optimizer = "bobyqa"))## boundary (singular) fit: see help('isSingular')Summary
## Linear mixed model fit by REML. t-tests use Satterthwaite's method [
## lmerModLmerTest]
## Formula: imp_height ~ political_orientation + (1 + political_orientation |
## country)
## Data: data_included_documented
## Control: lmerControl(optimizer = "bobyqa")
##
## REML criterion at convergence: 46915.5
##
## Scaled residuals:
## Min 1Q Median 3Q Max
## -3.2633 -0.5505 0.1130 0.7243 2.0751
##
## Random effects:
## Groups Name Variance Std.Dev. Corr
## country (Intercept) 8.972e-02 0.29953
## political_orientation 8.986e-05 0.00948 -1.00
## Residual 2.130e+00 1.45961
## Number of obs: 13026, groups: country, 143
##
## Fixed effects:
## Estimate Std. Error df t value Pr(>|t|)
## (Intercept) 3.779e+00 5.174e-02 5.157e+01 73.03 <2e-16 ***
## political_orientation 1.185e-01 9.667e-03 5.975e+02 12.26 <2e-16 ***
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
##
## Correlation of Fixed Effects:
## (Intr)
## pltcl_rnttn -0.597
## optimizer (bobyqa) convergence code: 0 (OK)
## boundary (singular) fit: see help('isSingular')## Computing profile confidence intervals ...## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): unexpected decrease in
## profile: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): unexpected decrease in
## profile: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): unexpected decrease in
## profile: using minstep
## Warning in nextpar(mat, cc, i, delta, lowcut, upcut): unexpected decrease in
## profile: using minstep## Warning in FUN(X[[i]], ...): non-monotonic profile for .sig01## Warning in FUN(X[[i]], ...): non-monotonic profile for .sig02## Warning in confint.thpr(pp, level = level, zeta = zeta): bad spline fit for
## .sig01: falling back to linear interpolation## Warning in confint.thpr(pp, level = level, zeta = zeta): bad spline fit for
## .sig02: falling back to linear interpolation## 0.15 % 99.85 %
## .sig01 0.18912373 0.39522206
## .sig02 -1.00000000 1.00000000
## .sig03 0.00000000 0.07675487
## .sigma 1.43715819 1.48624262
## (Intercept) 3.62468680 3.95869154
## political_orientation 0.08311549 0.15306504Standardized Coefficients
## # A tibble: 2 × 5
## Parameter Std_Coefficient CI CI_low CI_high
## <chr> <dbl> <dbl> <dbl> <dbl>
## 1 (Intercept) 0 0.997 0 0
## 2 political_orientation 0.108 0.997 0.0821 0.135H3b(2) Level Ratings for Partner’s Height
H3b(2) There is a positive linear link between right-wing political orientation and ideal partner’s height. Outcome: Level ratings for ideal partner’s height. Predictor: Political Orientation. Random intercept and random slope for country.
Models
model_ideal_height <- lmer(ideal_height ~ political_orientation + (1+political_orientation|country), data = data_included_documented, control =lmerControl(optimizer = "bobyqa"))## boundary (singular) fit: see help('isSingular')Summary
## Linear mixed model fit by REML. t-tests use Satterthwaite's method [
## lmerModLmerTest]
## Formula: ideal_height ~ political_orientation + (1 + political_orientation |
## country)
## Data: data_included_documented
## Control: lmerControl(optimizer = "bobyqa")
##
## REML criterion at convergence: 14686.1
##
## Scaled residuals:
## Min 1Q Median 3Q Max
## -7.0913 -0.1224 -0.0640 -0.0069 2.5128
##
## Random effects:
## Groups Name Variance Std.Dev. Corr
## country (Intercept) 1.611e-03 0.040138
## political_orientation 1.072e-05 0.003274 1.00
## Residual 1.882e-01 0.433773
## Number of obs: 12524, groups: country, 142
##
## Fixed effects:
## Estimate Std. Error df t value Pr(>|t|)
## (Intercept) 1.029e+00 1.091e-02 3.506e+01 94.284 <2e-16 ***
## political_orientation 3.100e-03 2.942e-03 3.668e+02 1.054 0.293
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
##
## Correlation of Fixed Effects:
## (Intr)
## pltcl_rnttn -0.556
## optimizer (bobyqa) convergence code: 0 (OK)
## boundary (singular) fit: see help('isSingular')## Computing profile confidence intervals ...## Warning in FUN(X[[i]], ...): non-monotonic profile for .sig02## Warning in confint.thpr(pp, level = level, zeta = zeta): bad spline fit for
## .sig02: falling back to linear interpolation## 0.15 % 99.85 %
## .sig01 0.010943539 0.08395169
## .sig02 -1.000000000 1.00000000
## .sig03 0.000000000 0.01532100
## .sigma 0.429230881 0.43981211
## (Intercept) 0.996323686 1.06969808
## political_orientation -0.007896281 0.01258552Standardized Coefficients
## # A tibble: 2 × 5
## Parameter Std_Coefficient CI CI_low CI_high
## <chr> <dbl> <dbl> <dbl> <dbl>
## 1 (Intercept) 0 0.997 0 0
## 2 political_orientation 0.00966 0.997 -0.0176 0.0369