Plots Effects of Contraception
Data
source("0_helpers.R")
library(extrafont)
theme_set(theme_tufte(base_size = 10, base_family='Arial'))
#plot specification for paper:
# apatheme=theme_bw()+
# theme(panel.grid.major=element_blank(),
# panel.grid.minor=element_blank(),
# panel.border=element_blank(),
# axis.line=element_line(),
# text = element_text(size=20),
# plot.title = element_text(size = 20))
load("data/cleaned_selected_wrangled.rdata")Selection Model
Hormonal Contraception
Simple Model
m_selection_hc_simple = brm(contraception_hormonal ~
age + net_income + relationship_duration_factor,
data = data,
family = bernoulli("probit"),
file = "m_selection_hc_simple")
m_selection_hc_simple_plot = m_selection_hc_simple %>%
spread_draws(b_age,
b_net_incomeeuro_500_1000, b_net_incomeeuro_1000_2000,
b_net_incomeeuro_2000_3000, b_net_incomeeuro_gt_3000, b_net_incomedont_tell,
b_relationship_duration_factorPartnered_upto12months,
b_relationship_duration_factorPartnered_upto28months,
b_relationship_duration_factorPartnered_upto52months,
b_relationship_duration_factorPartnered_morethan52months) %>%
pivot_longer(cols = c(b_age,
b_net_incomeeuro_500_1000, b_net_incomeeuro_1000_2000,
b_net_incomeeuro_2000_3000, b_net_incomeeuro_gt_3000, b_net_incomedont_tell,
b_relationship_duration_factorPartnered_upto12months,
b_relationship_duration_factorPartnered_upto28months,
b_relationship_duration_factorPartnered_upto52months,
b_relationship_duration_factorPartnered_morethan52months),
names_to = "condition",
values_to = "r_condition") %>%
mutate(condition_mean = r_condition,
group = ifelse(condition %contains% "b_relationship_duration_factor",
"Relationship Duration",
ifelse(condition %contains% "b_net_income",
"Income",
NA)),
condition = ifelse(condition == "b_age", "Age",
ifelse(condition == "b_net_incomeeuro_500_1000", "500 \U2013 1,000 Euro monthly",
ifelse(condition == "b_net_incomeeuro_1000_2000", "1,000 \U2013 2,000 Euro monthly",
ifelse(condition == "b_net_incomeeuro_2000_3000", "2,000 \U2013 3,000 Euro monthly",
ifelse(condition == "b_net_incomeeuro_gt_3000", ">3,000 Euro monthly",
ifelse(condition == "b_net_incomedont_tell", "do not disclose",
ifelse(condition == "b_relationship_duration_factorPartnered_upto12months",
"0 \U2013 12 months",
ifelse(condition == "b_relationship_duration_factorPartnered_upto28months",
"13 \U2013 28 months",
ifelse(condition == "b_relationship_duration_factorPartnered_upto52months",
"29 \U2013 52 months",
ifelse(condition == "b_relationship_duration_factorPartnered_morethan52months",
">52 months",
condition)))))))))),
group = ifelse(is.na(group), condition, group),
condition = factor(condition, levels = rev(c("Age",
"500 \U2013 1,000 Euro monthly", "1,000 \U2013 2,000 Euro monthly",
"2,000 \U2013 3,000 Euro monthly", ">3,000 Euro monthly", "do not disclose",
"0 \U2013 12 months", "13 \U2013 28 months", "29 \U2013 52 months",
">52 months")))) %>%
ggplot(aes(y = condition, x = condition_mean, color = group)) +
stat_halfeye(.width = 0.9) +
geom_vline(xintercept = 0, linetype = "dotted", size = 1) +
apatheme +
theme(legend.title = element_blank()) +
labs(x = "Effect Size Estimates", y = "Predictors") +
scale_x_continuous(limits = c(-1.2, 1.9),
breaks= c(-1.0, -0.5, 0, 0.5, 1.0, 1.5)) +
scale_color_manual(values = c("#1B9E77", "#D95F02", "#7570B3")) +
guides(color = guide_legend(override.aes = list(size = 10))) +
ggtitle(expression(paste(bold("Simple Model ("),
bolditalic("n"),
bold(" = 1,179)")))) +
theme(plot.title = element_text(hjust = -0.32))
m_selection_hc_simple_plot
Complex Model
m_selection_hc_complex = brm(contraception_hormonal ~
age + net_income + relationship_duration_factor +
education_years +
bfi_extra + bfi_neuro + bfi_agree + bfi_consc + bfi_open +
religiosity,
data = data,
family = bernoulli("probit"),
file = "m_selection_hc_complex")
m_selection_hc_complex_plot = m_selection_hc_complex %>%
spread_draws(b_age,
b_net_incomeeuro_500_1000, b_net_incomeeuro_1000_2000,
b_net_incomeeuro_2000_3000, b_net_incomeeuro_gt_3000, b_net_incomedont_tell,
b_relationship_duration_factorPartnered_upto12months,
b_relationship_duration_factorPartnered_upto28months,
b_relationship_duration_factorPartnered_upto52months,
b_relationship_duration_factorPartnered_morethan52months,
b_education_years,
b_bfi_extra, b_bfi_neuro, b_bfi_agree, b_bfi_consc, b_bfi_open,
b_religiosity) %>%
pivot_longer(cols = c(b_age,
b_net_incomeeuro_500_1000, b_net_incomeeuro_1000_2000,
b_net_incomeeuro_2000_3000, b_net_incomeeuro_gt_3000, b_net_incomedont_tell,
b_relationship_duration_factorPartnered_upto12months,
b_relationship_duration_factorPartnered_upto28months,
b_relationship_duration_factorPartnered_upto52months,
b_relationship_duration_factorPartnered_morethan52months,
b_education_years,
b_bfi_extra, b_bfi_neuro, b_bfi_agree, b_bfi_consc, b_bfi_open,
b_religiosity),
names_to = "condition",
values_to = "r_condition") %>%
mutate(condition_mean = r_condition,
group = ifelse(condition %contains% "b_relationship_duration_factor",
"Relationship Duration",
ifelse(condition %contains% "b_net_income",
"Income",
NA)),
condition = ifelse(condition == "b_age", "Age",
ifelse(condition == "b_net_incomeeuro_500_1000", "500 \U2013 1,000 Euro monthly",
ifelse(condition == "b_net_incomeeuro_1000_2000", "1,000 \U2013 2,000 Euro monthly",
ifelse(condition == "b_net_incomeeuro_2000_3000", "2,000 \U2013 3,000 Euro monthly",
ifelse(condition == "b_net_incomeeuro_gt_3000", ">3,000 Euro monthly",
ifelse(condition == "b_net_incomedont_tell", "do not disclose",
ifelse(condition == "b_relationship_duration_factorPartnered_upto12months",
"0 \U2013 12 months",
ifelse(condition == "b_relationship_duration_factorPartnered_upto28months",
"13 \U2013 28 months",
ifelse(condition == "b_relationship_duration_factorPartnered_upto52months",
"29 \U2013 52 months",
ifelse(condition == "b_relationship_duration_factorPartnered_morethan52months",
">52 months",
ifelse(condition == "b_education_years", "Years of Education",
ifelse(condition == "b_bfi_extra", "Extraversion",
ifelse(condition == "b_bfi_neuro", "Neuroticism",
ifelse(condition == "b_bfi_agree", "Agreeableness",
ifelse(condition == "b_bfi_consc", "Conscientiousness",
ifelse(condition == "b_bfi_open", "Openness",
ifelse(condition == "b_religiosity", "Religiosity",
condition))))))))))))))))),
group = ifelse(is.na(group), condition, group),
condition = factor(condition, levels = rev(c("Age",
"500 \U2013 1,000 Euro monthly", "1,000 \U2013 2,000 Euro monthly",
"2,000 \U2013 3,000 Euro monthly", ">3,000 Euro monthly", "do not disclose",
"0 \U2013 12 months", "13 \U2013 28 months", "29 \U2013 52 months",
">52 months",
"Years of Education",
"Extraversion", "Neuroticism", "Agreeableness",
"Conscientiousness","Openness","Religiosity"))),
group = factor(group, levels = c("Age", "Income", "Relationship Duration",
"Years of Education",
"Extraversion", "Neuroticism", "Agreeableness",
"Conscientiousness","Openness", "Religiosity"))) %>%
ggplot(aes(y = condition, x = condition_mean, color = group)) +
stat_halfeye(.width = 0.9) +
geom_vline(xintercept = 0, linetype = "dotted", size = 1) +
apatheme +
theme(legend.title = element_blank()) +
labs(x = "Effect Size Estimates", y = "Predictors") +
scale_x_continuous(limits = c(-1.2, 1.9),
breaks = c(-1.0, -0.5, 0, 0.5, 1.0, 1.5)) +
scale_color_manual(values = c("#1B9E77", "#D95F02", "#7570B3", "#E7298A", "#66A61E",
"#E6AB02", "#A6761D", "#0072B2", "#CC79A7", "#56B4E9")) +
guides(color = guide_legend(override.aes = list(size = 10))) +
ggtitle(expression(paste(bold("Complex Model ("),
bolditalic("n"),
bold(" = 1,179)")))) +
theme(plot.title = element_text(hjust = -0.32))
m_selection_hc_complex_plot
Combine Plots
selection_hc =
ggarrange(m_selection_hc_simple_plot + theme(legend.position = c(0.85,0.5),
plot.margin = margin(1,0,0,0,"cm")),
m_selection_hc_complex_plot + theme(legend.position = c(0.85,0.5),
plot.margin = margin(1,0,0,0,"cm")),
hjust = -0.1,
ncol = 1, nrow = 2,
align = "v")
selection_hc
jpeg('Effect size estimates for models with current contraceptive use as an outcome.jpg',
width = 900, height = 1000, quality = 100)
selection_hc
dev.off()## png
## 2(In)congruent use of hormonal contraceptives
Simple Model
m_selection_congruent_simple_onceHC = brm(congruent_contraception ~
age + net_income + relationship_duration_factor,
data = data %>% filter(contraception_meeting_partner == "yes"),
family = bernoulli("probit"),
file = "m_selection_congruent_simple_onceHC")
m_selection_congruent_simple_oncenonHC = brm(congruent_contraception ~
age + net_income + relationship_duration_factor,
data = data %>% filter(contraception_meeting_partner == "no"),
family = bernoulli("probit"),
file = "m_selection_congruent_simple_oncenonHC")
m_selection_congruent_simple_onceHC_plot = m_selection_congruent_simple_onceHC %>%
spread_draws(b_age,
b_net_incomeeuro_500_1000, b_net_incomeeuro_1000_2000,
b_net_incomeeuro_2000_3000, b_net_incomeeuro_gt_3000, b_net_incomedont_tell,
b_relationship_duration_factorPartnered_upto28months,
b_relationship_duration_factorPartnered_upto52months,
b_relationship_duration_factorPartnered_morethan52months) %>%
pivot_longer(cols = c(b_age,
b_net_incomeeuro_500_1000, b_net_incomeeuro_1000_2000,
b_net_incomeeuro_2000_3000, b_net_incomeeuro_gt_3000, b_net_incomedont_tell,
b_relationship_duration_factorPartnered_upto28months,
b_relationship_duration_factorPartnered_upto52months,
b_relationship_duration_factorPartnered_morethan52months),
names_to = "condition",
values_to = "r_condition") %>%
mutate(condition_mean = r_condition,
group = ifelse(condition %contains% "b_relationship_duration_factor",
"Relationship Duration",
ifelse(condition %contains% "b_net_income",
"Income",
NA)),
condition = ifelse(condition == "b_age", "Age",
ifelse(condition == "b_net_incomeeuro_500_1000", "500 \U2013 1,000 Euro monthly",
ifelse(condition == "b_net_incomeeuro_1000_2000", "1,000 \U2013 2,000 Euro monthly",
ifelse(condition == "b_net_incomeeuro_2000_3000", "2,000 \U2013 3,000 Euro monthly",
ifelse(condition == "b_net_incomeeuro_gt_3000", ">3,000 Euro monthly",
ifelse(condition == "b_net_incomedont_tell", "do not disclose",
ifelse(condition == "b_relationship_duration_factorPartnered_upto12months",
"0 \U2013 12 months",
ifelse(condition == "b_relationship_duration_factorPartnered_upto28months",
"13 \U2013 28 months",
ifelse(condition == "b_relationship_duration_factorPartnered_upto52months",
"29 \U2013 52 months",
ifelse(condition == "b_relationship_duration_factorPartnered_morethan52months",
">52 months",
condition)))))))))),
group = ifelse(is.na(group), condition, group),
condition = factor(condition, levels = rev(c("Age",
"500 \U2013 1,000 Euro monthly", "1,000 \U2013 2,000 Euro monthly",
"2,000 \U2013 3,000 Euro monthly", ">3,000 Euro monthly", "do not disclose",
"13 \U2013 28 months", "29 \U2013 52 months",
">52 months")))) %>%
ggplot(aes(y = condition, x = condition_mean, color = group)) +
stat_halfeye(.width = .90) +
geom_vline(xintercept = 0, linetype = "dotted", size = 1) +
apatheme +
theme(legend.title = element_blank()) +
labs(x = "Effect Size Estimates", y = "Predictors") +
scale_x_continuous(limits = c(-3, 3),
breaks= c(-3:3)) +
scale_color_manual(values = c("#1B9E77", "#D95F02", "#7570B3")) +
guides(color = guide_legend(override.aes = list(size = 5)))+
ggtitle(expression(paste(bold("Simple Model Initially HC User ("),
bolditalic("n"),
bold(" = 390)")))) +
theme(plot.title = element_text(hjust = -0.35))
m_selection_congruent_simple_onceHC_plot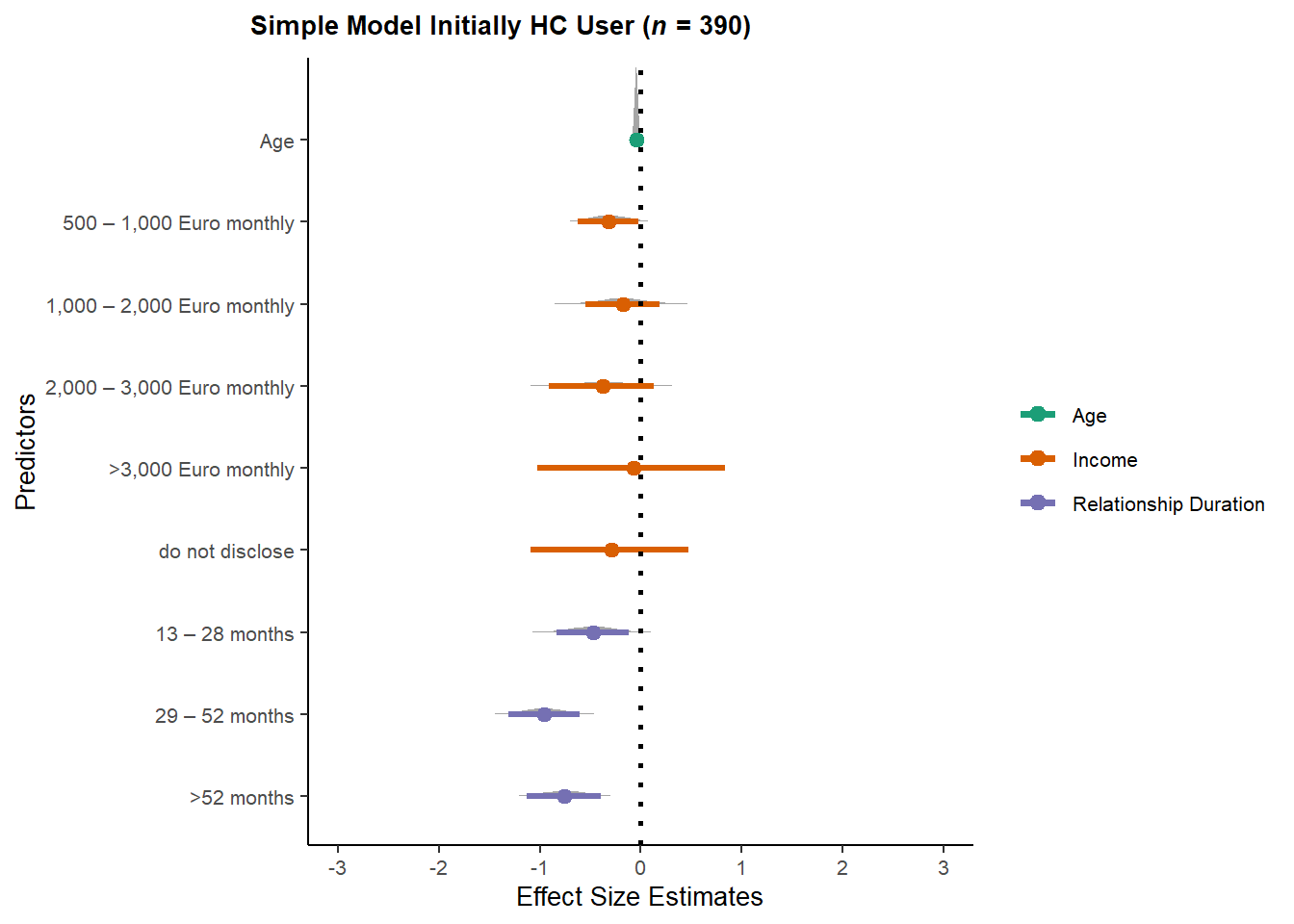
m_selection_congruent_simple_oncenonHC_plot = m_selection_congruent_simple_oncenonHC %>%
spread_draws(b_age,
b_net_incomeeuro_500_1000, b_net_incomeeuro_1000_2000,
b_net_incomeeuro_2000_3000, b_net_incomeeuro_gt_3000, b_net_incomedont_tell,
b_relationship_duration_factorPartnered_upto28months,
b_relationship_duration_factorPartnered_upto52months,
b_relationship_duration_factorPartnered_morethan52months) %>%
pivot_longer(cols = c(b_age,
b_net_incomeeuro_500_1000, b_net_incomeeuro_1000_2000,
b_net_incomeeuro_2000_3000, b_net_incomeeuro_gt_3000, b_net_incomedont_tell,
b_relationship_duration_factorPartnered_upto28months,
b_relationship_duration_factorPartnered_upto52months,
b_relationship_duration_factorPartnered_morethan52months),
names_to = "condition",
values_to = "r_condition") %>%
mutate(condition_mean = r_condition,
group = ifelse(condition %contains% "b_relationship_duration_factor",
"Relationship Duration",
ifelse(condition %contains% "b_net_income",
"Income",
NA)),
condition = ifelse(condition == "b_age", "Age",
ifelse(condition == "b_net_incomeeuro_500_1000", "500 \U2013 1,000 Euro monthly",
ifelse(condition == "b_net_incomeeuro_1000_2000", "1,000 \U2013 2,000 Euro monthly",
ifelse(condition == "b_net_incomeeuro_2000_3000", "2,000 \U2013 3,000 Euro monthly",
ifelse(condition == "b_net_incomeeuro_gt_3000", ">3,000 Euro monthly",
ifelse(condition == "b_net_incomedont_tell", "do not disclose",
ifelse(condition == "b_relationship_duration_factorPartnered_upto12months",
"0 \U2013 12 months",
ifelse(condition == "b_relationship_duration_factorPartnered_upto28months",
"13 \U2013 28 months",
ifelse(condition == "b_relationship_duration_factorPartnered_upto52months",
"29 \U2013 52 months",
ifelse(condition == "b_relationship_duration_factorPartnered_morethan52months",
">52 months",
condition)))))))))),
group = ifelse(is.na(group), condition, group),
condition = factor(condition, levels = rev(c("Age",
"500 \U2013 1,000 Euro monthly", "1,000 \U2013 2,000 Euro monthly",
"2,000 \U2013 3,000 Euro monthly", ">3,000 Euro monthly", "do not disclose",
"13 \U2013 28 months", "29 \U2013 52 months",
">52 months")))) %>%
ggplot(aes(y = condition, x = condition_mean, color = group)) +
stat_halfeye(.width = .90) +
geom_vline(xintercept = 0, linetype = "dotted", size = 1) +
apatheme +
theme(legend.title = element_blank()) +
labs(x = "Effect Size Estimates", y = "Predictors") +
scale_x_continuous(limits = c(-3, 3),
breaks= c(-3:3)) +
scale_color_manual(values = c("#1B9E77", "#D95F02", "#7570B3")) +
guides(color = guide_legend(override.aes = list(size = 5))) +
ggtitle(expression(paste(bold("Simple Model Initially Non-HC User ("),
bolditalic("n"),
bold(" = 384)")))) +
theme(plot.title = element_text(hjust = -0.35))
m_selection_congruent_simple_oncenonHC_plot
Complex Model
m_selection_congruent_complex_onceHC = brm(congruent_contraception ~
age + net_income + relationship_duration_factor +
education_years +
bfi_extra + bfi_neuro + bfi_agree + bfi_consc + bfi_open +
religiosity,
data = data %>% filter(contraception_meeting_partner == "yes"),
family = bernoulli("probit"),
file = "m_selection_congruent_complex_onceHC")
m_selection_congruent_complex_oncenonHC = brm(congruent_contraception ~
age + net_income + relationship_duration_factor +
education_years +
bfi_extra + bfi_neuro + bfi_agree + bfi_consc + bfi_open +
religiosity,
data = data %>% filter(contraception_meeting_partner == "no"),
family = bernoulli("probit"),
file = "m_selection_congruent_complex_oncenonHC")
m_selection_congruent_complex_onceHC_plot = m_selection_congruent_complex_onceHC %>%
spread_draws(b_age,
b_net_incomeeuro_500_1000, b_net_incomeeuro_1000_2000,
b_net_incomeeuro_2000_3000, b_net_incomeeuro_gt_3000, b_net_incomedont_tell,
b_relationship_duration_factorPartnered_upto28months,
b_relationship_duration_factorPartnered_upto52months,
b_relationship_duration_factorPartnered_morethan52months,
b_education_years,
b_bfi_extra, b_bfi_neuro, b_bfi_agree, b_bfi_consc, b_bfi_open,
b_religiosity) %>%
pivot_longer(cols = c(b_age,
b_net_incomeeuro_500_1000, b_net_incomeeuro_1000_2000,
b_net_incomeeuro_2000_3000, b_net_incomeeuro_gt_3000, b_net_incomedont_tell,
b_relationship_duration_factorPartnered_upto28months,
b_relationship_duration_factorPartnered_upto52months,
b_relationship_duration_factorPartnered_morethan52months,
b_education_years,
b_bfi_extra, b_bfi_neuro, b_bfi_agree, b_bfi_consc, b_bfi_open,
b_religiosity),
names_to = "condition",
values_to = "r_condition") %>%
mutate(condition_mean = r_condition,
group = ifelse(condition %contains% "b_relationship_duration_factor",
"Relationship Duration",
ifelse(condition %contains% "b_net_income",
"Income",
NA)),
condition = ifelse(condition == "b_age", "Age",
ifelse(condition == "b_net_incomeeuro_500_1000", "500 \U2013 1,000 Euro monthly",
ifelse(condition == "b_net_incomeeuro_1000_2000", "1,000 \U2013 2,000 Euro monthly",
ifelse(condition == "b_net_incomeeuro_2000_3000", "2,000 \U2013 3,000 Euro monthly",
ifelse(condition == "b_net_incomeeuro_gt_3000", ">3,000 Euro monthly",
ifelse(condition == "b_net_incomedont_tell", "do not disclose",
ifelse(condition == "b_relationship_duration_factorPartnered_upto12months",
"0 \U2013 12 months",
ifelse(condition == "b_relationship_duration_factorPartnered_upto28months",
"13 \U2013 28 months",
ifelse(condition == "b_relationship_duration_factorPartnered_upto52months",
"29 \U2013 52 months",
ifelse(condition == "b_relationship_duration_factorPartnered_morethan52months",
">52 months",
ifelse(condition == "b_education_years", "Years of Education",
ifelse(condition == "b_bfi_extra", "Extraversion",
ifelse(condition == "b_bfi_neuro", "Neuroticism",
ifelse(condition == "b_bfi_agree", "Agreeableness",
ifelse(condition == "b_bfi_consc", "Conscientiousness",
ifelse(condition == "b_bfi_open", "Openness",
ifelse(condition == "b_religiosity", "Religiosity",
condition))))))))))))))))),
group = ifelse(is.na(group), condition, group),
condition = factor(condition, levels = rev(c("Age",
"500 \U2013 1,000 Euro monthly", "1,000 \U2013 2,000 Euro monthly",
"2,000 \U2013 3,000 Euro monthly", ">3,000 Euro monthly", "do not disclose",
"13 \U2013 28 months", "29 \U2013 52 months",
">52 months",
"Years of Education",
"Extraversion", "Neuroticism", "Agreeableness",
"Conscientiousness","Openness","Religiosity"))),
group = factor(group, levels = c("Age", "Income", "Relationship Duration",
"Years of Education",
"Extraversion", "Neuroticism", "Agreeableness",
"Conscientiousness","Openness","Religiosity"))) %>%
ggplot(aes(y = condition, x = condition_mean, color = group)) +
stat_halfeye(.width = .90) +
geom_vline(xintercept = 0, linetype = "dotted", size = 1) +
apatheme +
theme(legend.title = element_blank()) +
labs(x = "Effect Size Estimates", y = "Predictors") +
scale_x_continuous(limits = c(-3, 3),
breaks = c(-3:3)) +
scale_color_manual(values = c("#1B9E77", "#D95F02", "#7570B3", "#E7298A", "#66A61E",
"#E6AB02", "#A6761D", "#0072B2", "#CC79A7", "#56B4E9")) +
guides(color = guide_legend(override.aes = list(size = 5)))+
ggtitle(expression(paste(bold("Complex Model Initially HC User ("),
bolditalic("n"),
bold(" = 390)")))) +
theme(plot.title = element_text(hjust = -0.35))
m_selection_congruent_complex_onceHC_plot
m_selection_congruent_complex_oncenonHC_plot = m_selection_congruent_complex_oncenonHC %>%
spread_draws(b_age,
b_net_incomeeuro_500_1000, b_net_incomeeuro_1000_2000,
b_net_incomeeuro_2000_3000, b_net_incomeeuro_gt_3000, b_net_incomedont_tell,
b_relationship_duration_factorPartnered_upto28months,
b_relationship_duration_factorPartnered_upto52months,
b_relationship_duration_factorPartnered_morethan52months,
b_education_years,
b_bfi_extra, b_bfi_neuro, b_bfi_agree, b_bfi_consc, b_bfi_open,
b_religiosity) %>%
pivot_longer(cols = c(b_age,
b_net_incomeeuro_500_1000, b_net_incomeeuro_1000_2000,
b_net_incomeeuro_2000_3000, b_net_incomeeuro_gt_3000, b_net_incomedont_tell,
b_relationship_duration_factorPartnered_upto28months,
b_relationship_duration_factorPartnered_upto52months,
b_relationship_duration_factorPartnered_morethan52months,
b_education_years,
b_bfi_extra, b_bfi_neuro, b_bfi_agree, b_bfi_consc, b_bfi_open,
b_religiosity),
names_to = "condition",
values_to = "r_condition") %>%
mutate(condition_mean = r_condition,
group = ifelse(condition %contains% "b_relationship_duration_factor",
"Relationship Duration",
ifelse(condition %contains% "b_net_income",
"Income",
NA)),
condition = ifelse(condition == "b_age", "Age",
ifelse(condition == "b_net_incomeeuro_500_1000", "500 \U2013 1,000 Euro monthly",
ifelse(condition == "b_net_incomeeuro_1000_2000", "1,000 \U2013 2,000 Euro monthly",
ifelse(condition == "b_net_incomeeuro_2000_3000", "2,000 \U2013 3,000 Euro monthly",
ifelse(condition == "b_net_incomeeuro_gt_3000", ">3,000 Euro monthly",
ifelse(condition == "b_net_incomedont_tell", "do not disclose",
ifelse(condition == "b_relationship_duration_factorPartnered_upto12months",
"0 \U2013 12 months",
ifelse(condition == "b_relationship_duration_factorPartnered_upto28months",
"13 \U2013 28 months",
ifelse(condition == "b_relationship_duration_factorPartnered_upto52months",
"29 \U2013 52 months",
ifelse(condition == "b_relationship_duration_factorPartnered_morethan52months",
">52 months",
ifelse(condition == "b_education_years", "Years of Education",
ifelse(condition == "b_bfi_extra", "Extraversion",
ifelse(condition == "b_bfi_neuro", "Neuroticism",
ifelse(condition == "b_bfi_agree", "Agreeableness",
ifelse(condition == "b_bfi_consc", "Conscientiousness",
ifelse(condition == "b_bfi_open", "Openness",
ifelse(condition == "b_religiosity", "Religiosity",
condition))))))))))))))))),
group = ifelse(is.na(group), condition, group),
condition = factor(condition, levels = rev(c("Age",
"500 \U2013 1,000 Euro monthly", "1,000 \U2013 2,000 Euro monthly",
"2,000 \U2013 3,000 Euro monthly", ">3,000 Euro monthly", "do not disclose",
"13 \U2013 28 months", "29 \U2013 52 months",
">52 months",
"Years of Education",
"Extraversion", "Neuroticism", "Agreeableness",
"Conscientiousness","Openness","Religiosity"))),
group = factor(group, levels = c("Age", "Income", "Relationship Duration",
"Years of Education",
"Extraversion", "Neuroticism", "Agreeableness",
"Conscientiousness","Openness","Religiosity"))) %>%
ggplot(aes(y = condition, x = condition_mean, color = group)) +
stat_halfeye(.width = .90) +
geom_vline(xintercept = 0, linetype = "dotted", size = 1) +
apatheme +
theme(legend.title = element_blank()) +
labs(x = "Effect Size Estimates", y = "Predictors") +
scale_x_continuous(limits = c(-3, 3),
breaks = c(-3:3)) +
scale_color_manual(values = c("#1B9E77", "#D95F02", "#7570B3", "#E7298A", "#66A61E",
"#E6AB02", "#A6761D", "#0072B2", "#CC79A7", "#56B4E9")) +
guides(color = guide_legend(override.aes = list(size = 5)))+
ggtitle(expression(paste(bold("Complex Model Initially Non-HC User ("),
bolditalic("n"),
bold(" = 384)")))) +
theme(plot.title = element_text(hjust = -0.35))
m_selection_congruent_complex_oncenonHC_plot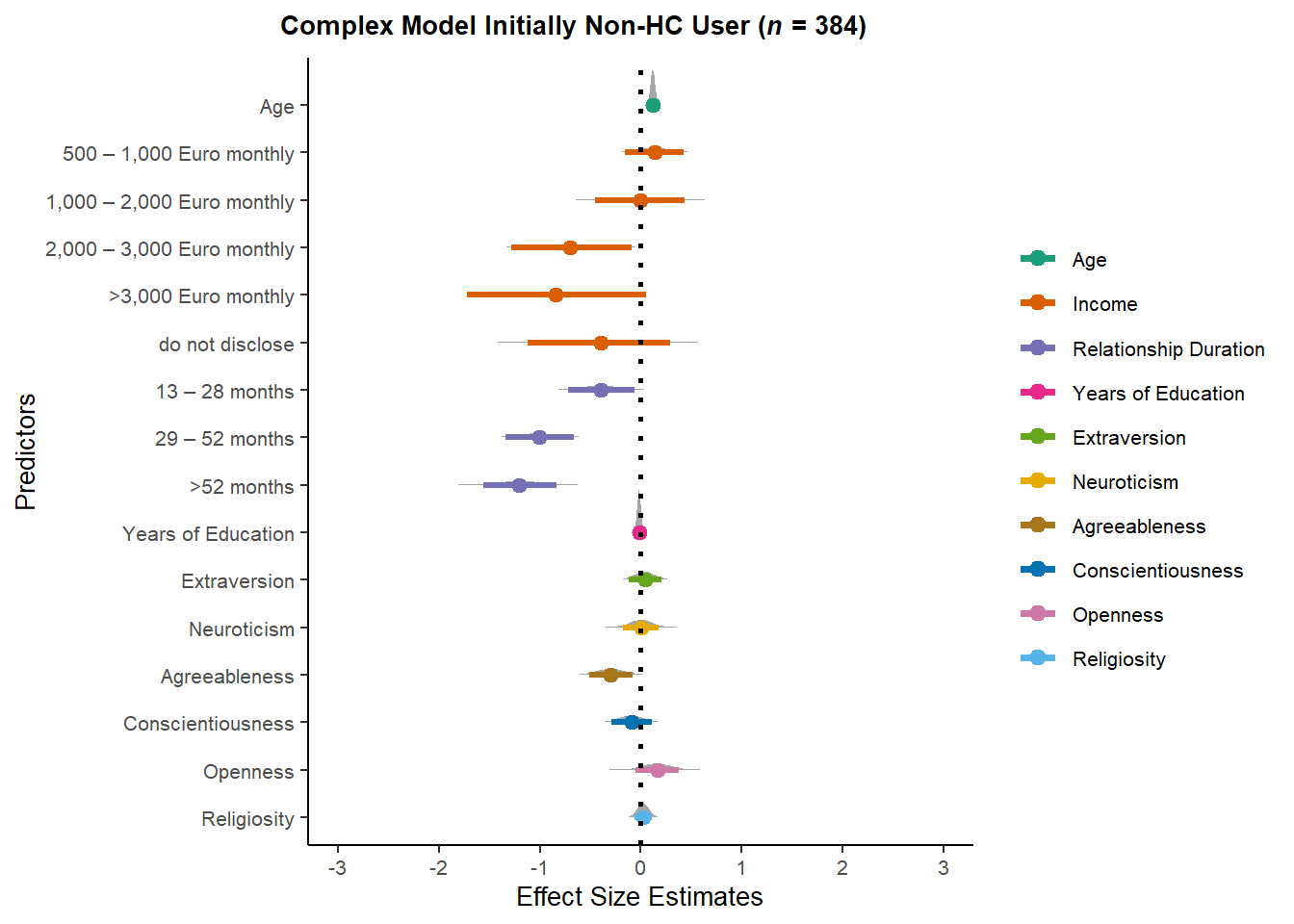
Combine Plots
selection_congruent =
ggarrange(m_selection_congruent_simple_onceHC_plot +
theme(legend.position = "none", plot.margin = margin(1,0,0,0,"cm")),
m_selection_congruent_simple_oncenonHC_plot +
theme(legend.position = c(0.85,0.5), plot.margin = margin(1,0,0,0,"cm")),
m_selection_congruent_complex_onceHC_plot +
theme(legend.position = "none", plot.margin = margin(1,0,0,0,"cm")),
m_selection_congruent_complex_oncenonHC_plot +
theme(legend.position = c(0.85,0.5), plot.margin = margin(1,0,0,0,"cm")),
hjust = -0.1, vjust = 1.1,
ncol = 2, nrow = 2,
align = "v")
selection_congruent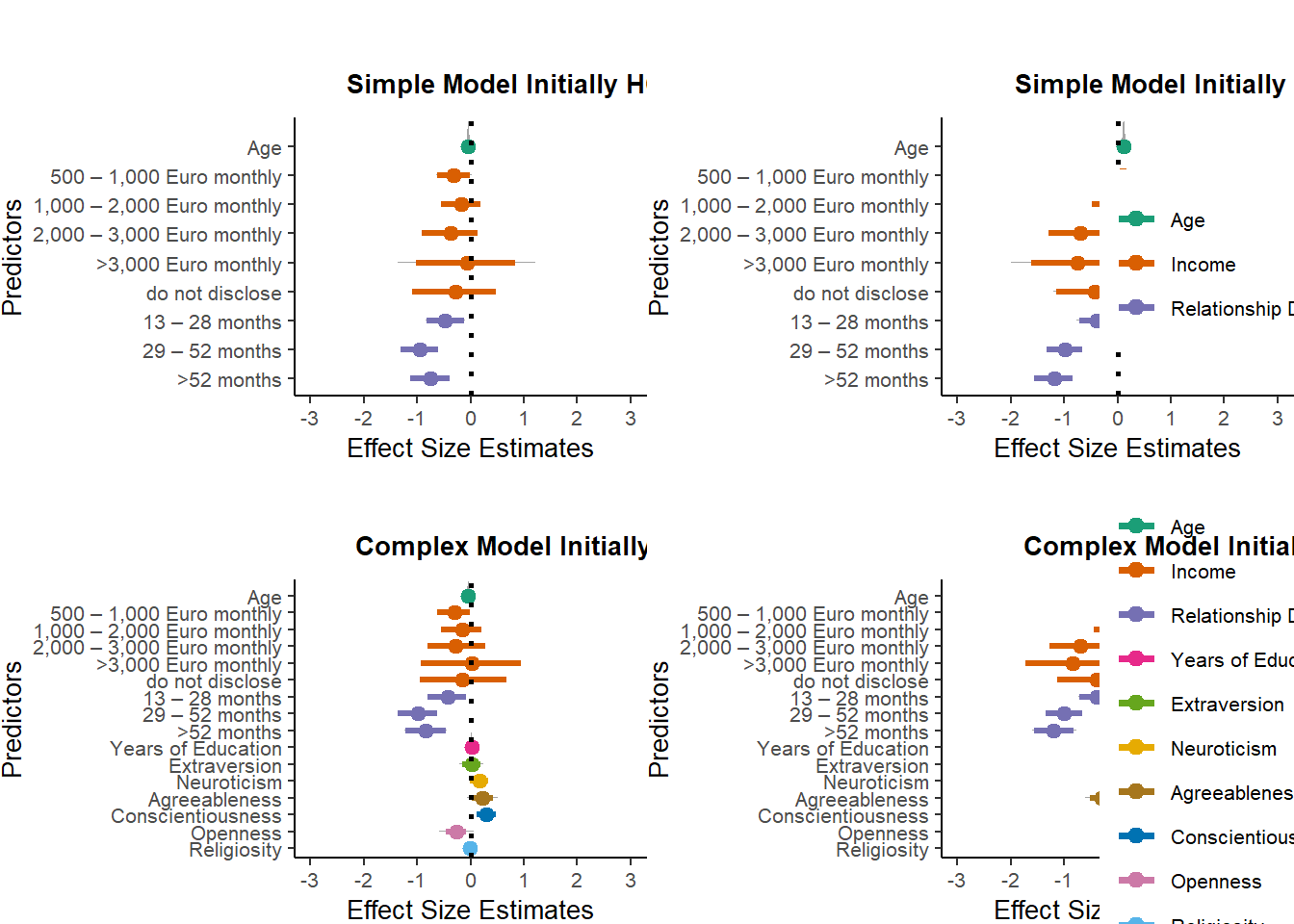
jpeg('Effect size estimates for models with congruency in contraceptive method as an outcome.jpg',
width = 1600, height = 720, quality = 100)
selection_congruent
dev.off()## png
## 2Effects of Hormonal Contraceptives
Uncontrolled
Models
m_hc_atrr = brm(attractiveness_partner ~ contraception_hormonal,
data = data, family = gaussian(),
file = "m_hc_atrr")
m_hc_relsat = brm(relationship_satisfaction ~ contraception_hormonal,
data = data, family = gaussian(),
file = "m_hc_relsat")
m_hc_sexsat = brm(satisfaction_sexual_intercourse ~ contraception_hormonal,
data = data, family = gaussian(),
file = "m_hc_sexsat")
m_hc_libido = brm(diary_libido_mean ~ contraception_hormonal,
data = data, family = gaussian(),
file = "m_hc_libido")
m_hc_sexfreqpen = brm(diary_sex_active_sex_sum ~
offset(log(number_of_days)) +
contraception_hormonal,
data = data, family = poisson(),
file = "mrobust_hc_sexfreqpen")
m_hc_masfreq = brm(diary_masturbation_sum ~ offset(log(number_of_days)) +
contraception_hormonal,
data = data, family = poisson(),
file = "mrobust_hc_masfreq")Forest Plot for Effect Sizes
models = rbind(
m_hc_atrr %>%
spread_draws(b_contraception_hormonalyes) %>%
pivot_longer(cols = c(b_contraception_hormonalyes),
names_to = "condition",
values_to = "r_condition") %>%
mutate(model = "Perceived Partner Attractiveness\nROPE = [-0.07, 0.07]",
ROPE = 0.07),
m_hc_relsat %>%
spread_draws(b_contraception_hormonalyes) %>%
pivot_longer(cols = c(b_contraception_hormonalyes),
names_to = "condition",
values_to = "r_condition") %>%
mutate(model = "Relationship Satisfaction\nROPE = [-0.04, 0.04]",
ROPE = 0.04),
m_hc_sexsat %>%
spread_draws(b_contraception_hormonalyes) %>%
pivot_longer(cols = c(b_contraception_hormonalyes),
names_to = "condition",
values_to = "r_condition") %>%
mutate(model = "Sexual Satisfaction\nROPE = [-0.11, 0.11]",
ROPE = 0.11),
m_hc_libido %>%
spread_draws(b_contraception_hormonalyes) %>%
pivot_longer(cols = c(b_contraception_hormonalyes),
names_to = "condition",
values_to = "r_condition") %>%
mutate(model = "Libido\nROPE = [-0.06, 0.06]",
ROPE = 0.06),
m_hc_sexfreqpen %>%
spread_draws(b_contraception_hormonalyes) %>%
pivot_longer(cols = c(b_contraception_hormonalyes),
names_to = "condition",
values_to = "r_condition") %>%
mutate(model = "Frequency of Vaginal Intercourse\nROPE = [-0.05, 0.05]",
ROPE = 0.05),
m_hc_masfreq %>%
spread_draws(b_contraception_hormonalyes) %>%
pivot_longer(cols = c(b_contraception_hormonalyes),
names_to = "condition",
values_to = "r_condition") %>%
mutate(model = "Frequency of Masturbation\nROPE = [-0.05, 0.05]",
ROPE = 0.05))
data1 = models %>%
mutate(model = factor(model,
levels = c("Frequency of Masturbation\nROPE = [-0.05, 0.05]",
"Frequency of Vaginal Intercourse\nROPE = [-0.05, 0.05]",
"Libido\nROPE = [-0.06, 0.06]",
"Sexual Satisfaction\nROPE = [-0.11, 0.11]",
"Relationship Satisfaction\nROPE = [-0.04, 0.04]",
"Perceived Partner Attractiveness\nROPE = [-0.07, 0.07]")))
data2 = models %>%
group_by(model, ROPE) %>%
mutate(model = factor(model,
levels = c("Frequency of Masturbation\nROPE = [-0.05, 0.05]",
"Frequency of Vaginal Intercourse\nROPE = [-0.05, 0.05]",
"Libido\nROPE = [-0.06, 0.06]",
"Sexual Satisfaction\nROPE = [-0.11, 0.11]",
"Relationship Satisfaction\nROPE = [-0.04, 0.04]",
"Perceived Partner Attractiveness\nROPE = [-0.07, 0.07]"))) %>%
mean_qi(condition_mean = r_condition,
.width = c(.90)) %>%
mutate(text = ifelse(.width == .90,
str_c(round(condition_mean, 2),
" [",
round(.lower, 2),
"; ",
round(.upper, 2),
"]"), NA),
sig = ifelse(.lower > ROPE | .upper < -ROPE, T, F))
hc_uncontrolled = ggplot() +
stat_halfeye(data = data1,
aes(y = model, x = r_condition,
color = "grey"),
.width = .90) +
geom_vline(xintercept = 0, linetype = "dotted") +
geom_pointinterval(data = data2,
aes(y = model,
x = condition_mean,
xmin = .lower,
xmax = .upper,
color = sig)) +
geom_point(data = data1, aes(y = model, x = ROPE), shape = "|", size = 10, color = "black") +
geom_point(data = data1, aes(y = model, x = -ROPE), shape = "|", size = 10, color = "black") +
geom_point(data = data1, aes(y = model, x = ROPE), shape = 58, size = 10, color = "white") +
geom_point(data = data1, aes(y = model, x = -ROPE), shape = 58, size = 10, color = "white") +
apatheme +
scale_color_manual(data2, values = c("skyblue", "grey", "red")) +
scale_x_continuous(breaks = c(-0.5, -0.4, -0.3, -0.2, -0.1, 0, 0.1, 0.2, 0.3, 0.4, 0.5),
limits = c(-0.5, 0.5)) +
labs(x = "Effect Size Estimates of Hormonal Contraception", y = "Outcomes") +
theme(legend.position = "none")
hc_uncontrolled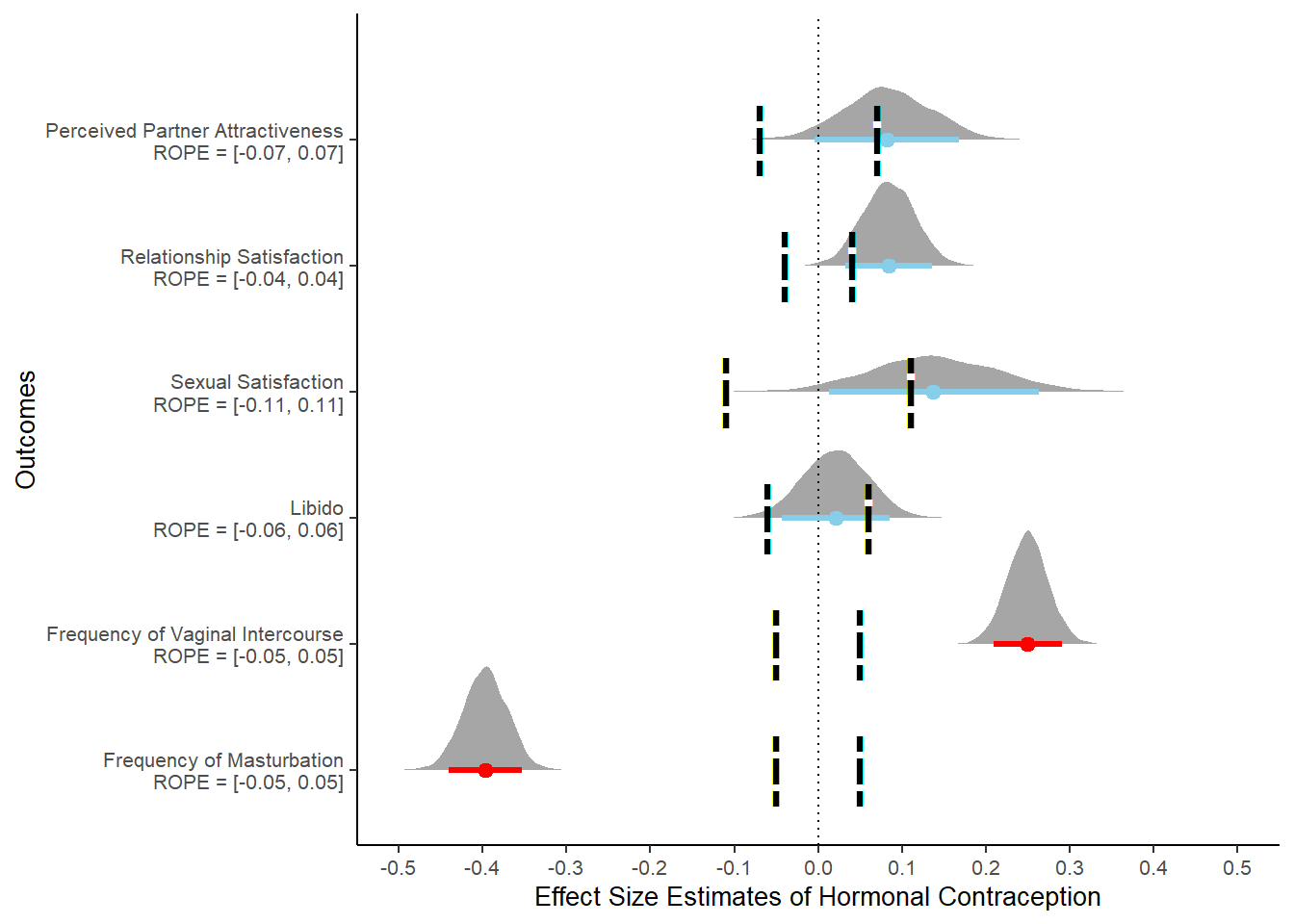
Controlled
Models
m_hc_atrr_controlled = brm(attractiveness_partner ~ contraception_hormonal +
age + net_income + relationship_duration_factor +
education_years +
bfi_extra + bfi_neuro + bfi_agree + bfi_consc + bfi_open +
religiosity,
data = data, family = gaussian(),
file = "m_hc_atrr_controlled")
m_hc_relsat_controlled = brm(relationship_satisfaction ~ contraception_hormonal +
age + net_income + relationship_duration_factor +
education_years +
bfi_extra + bfi_neuro + bfi_agree + bfi_consc + bfi_open +
religiosity,
data = data, family = gaussian(),
file = "m_hc_relsat_controlled")
m_hc_sexsat_controlled = brm(satisfaction_sexual_intercourse ~ contraception_hormonal +
age + net_income + relationship_duration_factor +
education_years +
bfi_extra + bfi_neuro + bfi_agree + bfi_consc + bfi_open +
religiosity,
data = data, family = gaussian(),
file = "m_hc_sexsat_controlled")
m_hc_libido_controlled = brm(diary_libido_mean ~ contraception_hormonal +
age + net_income + relationship_duration_factor +
education_years +
bfi_extra + bfi_neuro + bfi_agree + bfi_consc + bfi_open +
religiosity,
data = data, family = gaussian(),
file = "m_hc_libido_controlled")
m_hc_sexfreqpen_controlled = brm(diary_sex_active_sex_sum ~
offset(log(number_of_days)) +
contraception_hormonal +
age + net_income + relationship_duration_factor +
education_years +
bfi_extra + bfi_neuro + bfi_agree + bfi_consc + bfi_open +
religiosity,
data = data, family = poisson(),
file = "mrobust_hc_sexfreqpen_controlled")
m_hc_masfreq_controlled = brm(diary_masturbation_sum ~
offset(log(number_of_days)) +
contraception_hormonal +
age + net_income + relationship_duration_factor +
education_years +
bfi_extra + bfi_neuro + bfi_agree + bfi_consc + bfi_open +
religiosity,
data = data, family = poisson(),
file = "mrobust_hc_masfreq_controlled")Forest Plot for Effect Sizes
models = rbind(
m_hc_atrr_controlled %>%
spread_draws(b_contraception_hormonalyes) %>%
pivot_longer(cols = c(b_contraception_hormonalyes),
names_to = "condition",
values_to = "r_condition") %>%
mutate(model = "Perceived Partner Attractiveness\nROPE = [-0.07, 0.07]",
ROPE = 0.07),
m_hc_relsat_controlled %>%
spread_draws(b_contraception_hormonalyes) %>%
pivot_longer(cols = c(b_contraception_hormonalyes),
names_to = "condition",
values_to = "r_condition") %>%
mutate(model = "Relationship Satisfaction\nROPE = [-0.04, 0.04]",
ROPE = 0.04),
m_hc_sexsat_controlled %>%
spread_draws(b_contraception_hormonalyes) %>%
pivot_longer(cols = c(b_contraception_hormonalyes),
names_to = "condition",
values_to = "r_condition") %>%
mutate(model = "Sexual Satisfaction\nROPE = [-0.11, 0.11]",
ROPE = 0.11),
m_hc_libido_controlled %>%
spread_draws(b_contraception_hormonalyes) %>%
pivot_longer(cols = c(b_contraception_hormonalyes),
names_to = "condition",
values_to = "r_condition") %>%
mutate(model = "Libido\nROPE = [-0.06, 0.06]",
ROPE = 0.06),
m_hc_sexfreqpen_controlled %>%
spread_draws(b_contraception_hormonalyes) %>%
pivot_longer(cols = c(b_contraception_hormonalyes),
names_to = "condition",
values_to = "r_condition") %>%
mutate(model = "Frequency of Vaginal Intercourse\nROPE = [-0.05, 0.05]",
ROPE = 0.05),
m_hc_masfreq_controlled %>%
spread_draws(b_contraception_hormonalyes) %>%
pivot_longer(cols = c(b_contraception_hormonalyes),
names_to = "condition",
values_to = "r_condition") %>%
mutate(model = "Frequency of Masturbation\nROPE = [-0.05, 0.05]",
ROPE = 0.05))
data1 = models %>%
mutate(model = factor(model,
levels = c("Frequency of Masturbation\nROPE = [-0.05, 0.05]",
"Frequency of Vaginal Intercourse\nROPE = [-0.05, 0.05]",
"Libido\nROPE = [-0.06, 0.06]",
"Sexual Satisfaction\nROPE = [-0.11, 0.11]",
"Relationship Satisfaction\nROPE = [-0.04, 0.04]",
"Perceived Partner Attractiveness\nROPE = [-0.07, 0.07]")))
data2 = models %>%
group_by(model, ROPE) %>%
mutate(model = factor(model,
levels = c("Frequency of Masturbation\nROPE = [-0.05, 0.05]",
"Frequency of Vaginal Intercourse\nROPE = [-0.05, 0.05]",
"Libido\nROPE = [-0.06, 0.06]",
"Sexual Satisfaction\nROPE = [-0.11, 0.11]",
"Relationship Satisfaction\nROPE = [-0.04, 0.04]",
"Perceived Partner Attractiveness\nROPE = [-0.07, 0.07]"))) %>%
mean_qi(condition_mean = r_condition,
.width = c(.90)) %>%
mutate(text = ifelse(.width == .90,
str_c(round(condition_mean, 2),
" [",
round(.lower, 2),
"; ",
round(.upper, 2),
"]"), NA),
sig = ifelse(.lower > ROPE | .upper < -ROPE, "declined",
ifelse(abs(.lower) < ROPE & .upper > ROPE, "undecidable", "accepted")))
hc_controlled = ggplot() +
stat_halfeye(data = data1,
aes(y = model, x = r_condition,
color = "grey"),
.width = .90) +
geom_vline(xintercept = 0, linetype = "dotted") +
geom_pointinterval(data = data2,
aes(y = model,
x = condition_mean,
xmin = .lower,
xmax = .upper,
color = sig)) +
geom_point(data = data1, aes(y = model, x = ROPE), shape = "|", size = 10, color = "black") +
geom_point(data = data1, aes(y = model, x = -ROPE), shape = "|", size = 10, color = "black") +
geom_point(data = data1, aes(y = model, x = ROPE), shape = 58, size = 10, color = "white") +
geom_point(data = data1, aes(y = model, x = -ROPE), shape = 58, size = 10, color = "white") +
apatheme +
theme(legend.position = "none") +
scale_color_manual(values = c("red", "green", "skyblue", "grey")) +
scale_x_continuous(breaks = c(-0.5, -0.4, -0.3, -0.2, -0.1, 0, 0.1, 0.2, 0.3, 0.4, 0.5),
limits = c(-0.5, 0.5)) +
labs(x = "Effect Size Estimates of Hormonal Contraception", y = "Outcomes") +
theme(legend.position = "none")
hc_controlled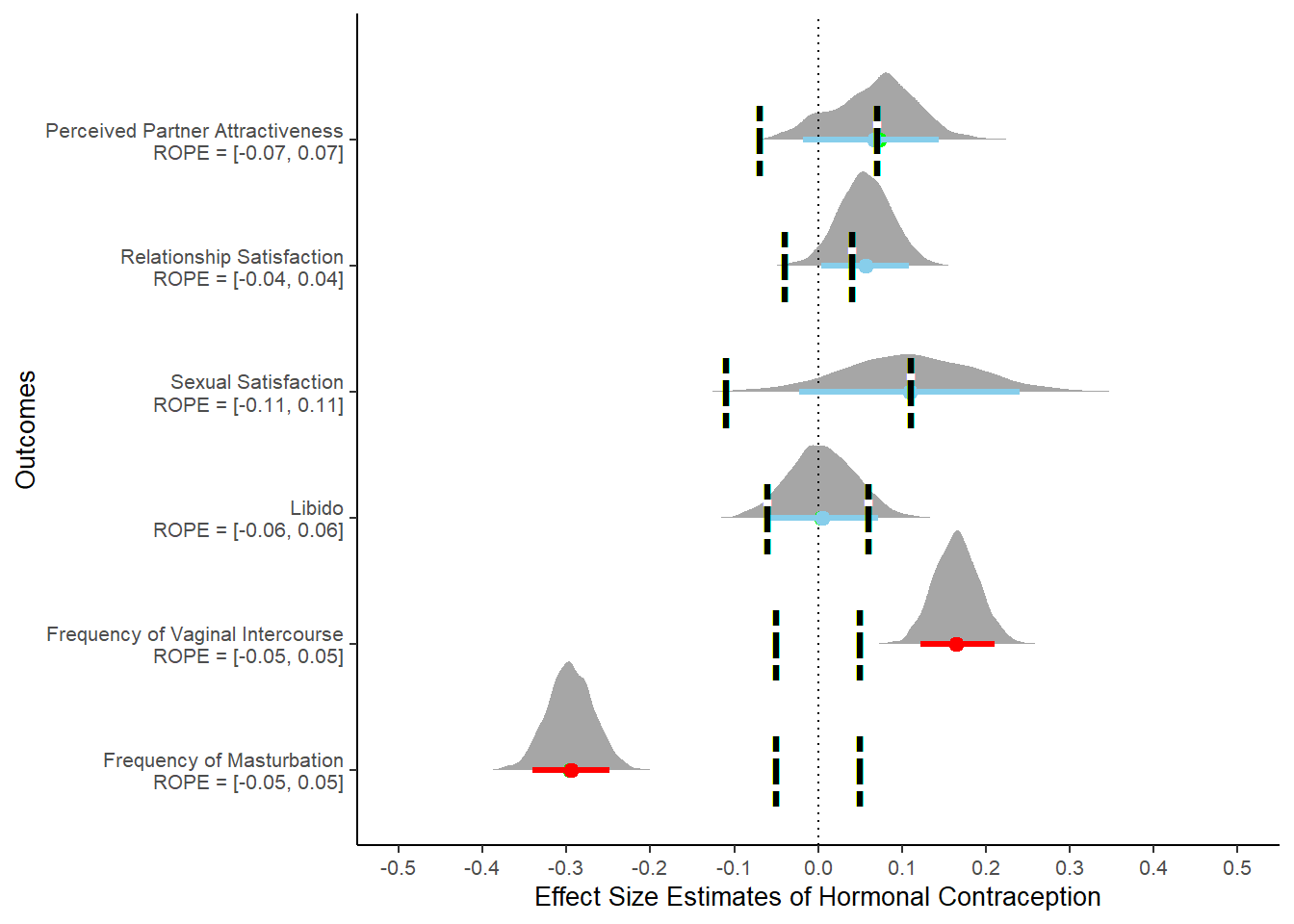
Combine Plots
hc_plots =
ggarrange(hc_uncontrolled + theme(plot.margin = margin(1,0,0,0,"cm")),
hc_controlled + theme(plot.margin = margin(3,0,0,0,"cm")),
labels = c("Uncontrolled Model", "\nControlled Model"),
font.label = list(size = 20, color = "black", face = "bold", family = "sans"),
hjust = -0.1,
ncol = 1, nrow = 2,
align = "v")
hc_plots
jpeg('Effect size estimates for hormonal contraceptives on outcomes.jpg',
width = 800, height = 1000, quality = 100)
hc_plots
dev.off()## png
## 2Effects of (in)congruent use of hormonal contraceptives
Plots
m_congruency_atrr = brm(attractiveness_partner ~
contraception_hormonal * congruent_contraception,
data = data, family = gaussian(),
file = "m_congruency_atrr")
m_congruency_atrr_ce = m_congruency_atrr %>%
conditional_effects(., effects = "contraception_hormonal:congruent_contraception",
prob = 0.90)
m_congruency_atrr_plot =
plot(m_congruency_atrr_ce, plot = FALSE)[[1]] +
labs(x = "Current Contraceptive Method",
y = "Perceived Partner Attractiveness") +
scale_x_discrete(labels=c("Non-HC", "HC")) +
scale_color_manual(name = "Congruent Contraceptive Method", labels = c("No", "Yes"),
values=c("#D95F02", "#1B9E77")) +
scale_fill_manual(name = "Congruent Contraceptive Method", labels = c("No", "Yes"),
values=c("#D95F02", "#1B9E77")) +
apatheme
m_congruency_atrr_plot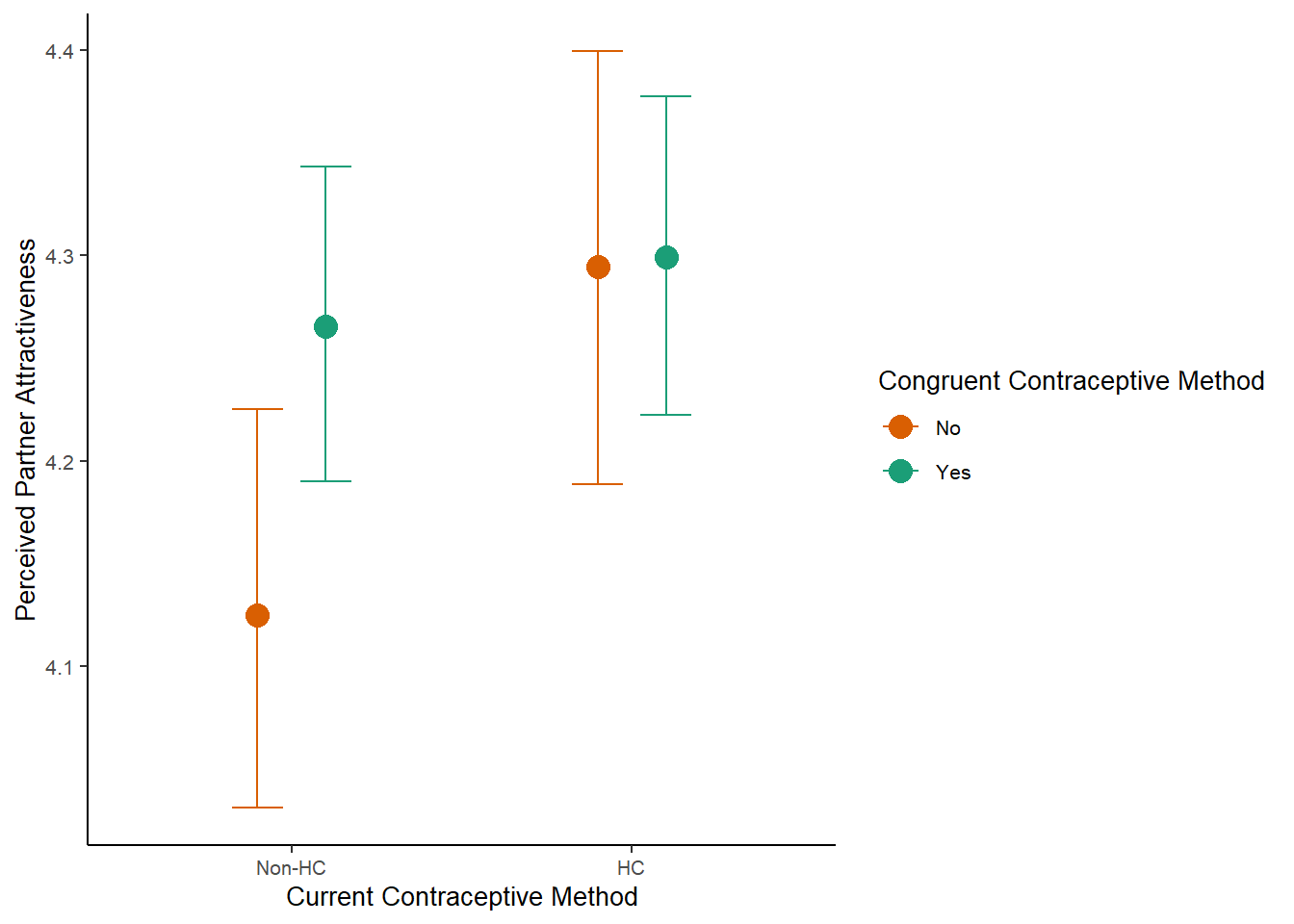
m_congruency_relsat = brm(relationship_satisfaction ~
contraception_hormonal * congruent_contraception,
data = data, family = gaussian(),
file = "m_congruency_relsat")
m_congruency_relsat_ce = m_congruency_relsat %>%
conditional_effects(., effects = "contraception_hormonal:congruent_contraception",
prob = 0.90)
m_congruency_relsat_plot =
plot(m_congruency_relsat_ce, plot = FALSE)[[1]] +
labs(x = "Current Contraceptive Method",
y = "Relationship Satisfaction") +
scale_x_discrete(labels=c("Non-HC", "HC")) +
scale_color_manual(name = "Congruent Contraceptive Method", labels = c("No", "Yes"),
values=c("#D95F02", "#1B9E77")) +
scale_fill_manual(name = "Congruent Contraceptive Method", labels = c("No", "Yes"),
values=c("#D95F02", "#1B9E77")) +
apatheme
m_congruency_relsat_plot
m_congruency_sexsat = brm(satisfaction_sexual_intercourse ~
contraception_hormonal * congruent_contraception,
data = data, family = gaussian(),
file = "m_congruency_sexsat")
m_congruency_sexsat_ce = m_congruency_sexsat %>%
conditional_effects(., effects = "contraception_hormonal:congruent_contraception",
prob = 0.90)
m_congruency_sexsat_plot =
plot(m_congruency_sexsat_ce, plot = FALSE)[[1]] +
labs(x = "Current Contraceptive Method",
y = "Sexual Satisfaction") +
scale_x_discrete(labels=c("Non-HC", "HC")) +
scale_color_manual(name = "Congruent Contraceptive Method", labels = c("No", "Yes"),
values=c("#D95F02", "#1B9E77")) +
scale_fill_manual(name = "Congruent Contraceptive Method", labels = c("No", "Yes"),
values=c("#D95F02", "#1B9E77")) +
apatheme
m_congruency_sexsat_plot
m_congruency_libido = brm(diary_libido_mean ~
contraception_hormonal * congruent_contraception,
data = data, family = gaussian(),
file = "m_congruency_libido")
m_congruency_libido_ce = m_congruency_libido %>%
conditional_effects(., effects = "contraception_hormonal:congruent_contraception",
prob = 0.90)
m_congruency_libido_plot =
plot(m_congruency_libido_ce, plot = FALSE)[[1]] +
labs(x = "Current Contraceptive Method",
y = "Libido") +
scale_x_discrete(labels=c("Non-HC", "HC")) +
scale_color_manual(name = "Congruent Contraceptive Method", labels = c("No", "Yes"),
values=c("#D95F02", "#1B9E77")) +
scale_fill_manual(name = "Congruent Contraceptive Method", labels = c("No", "Yes"),
values=c("#D95F02", "#1B9E77")) +
apatheme
m_congruency_libido_plot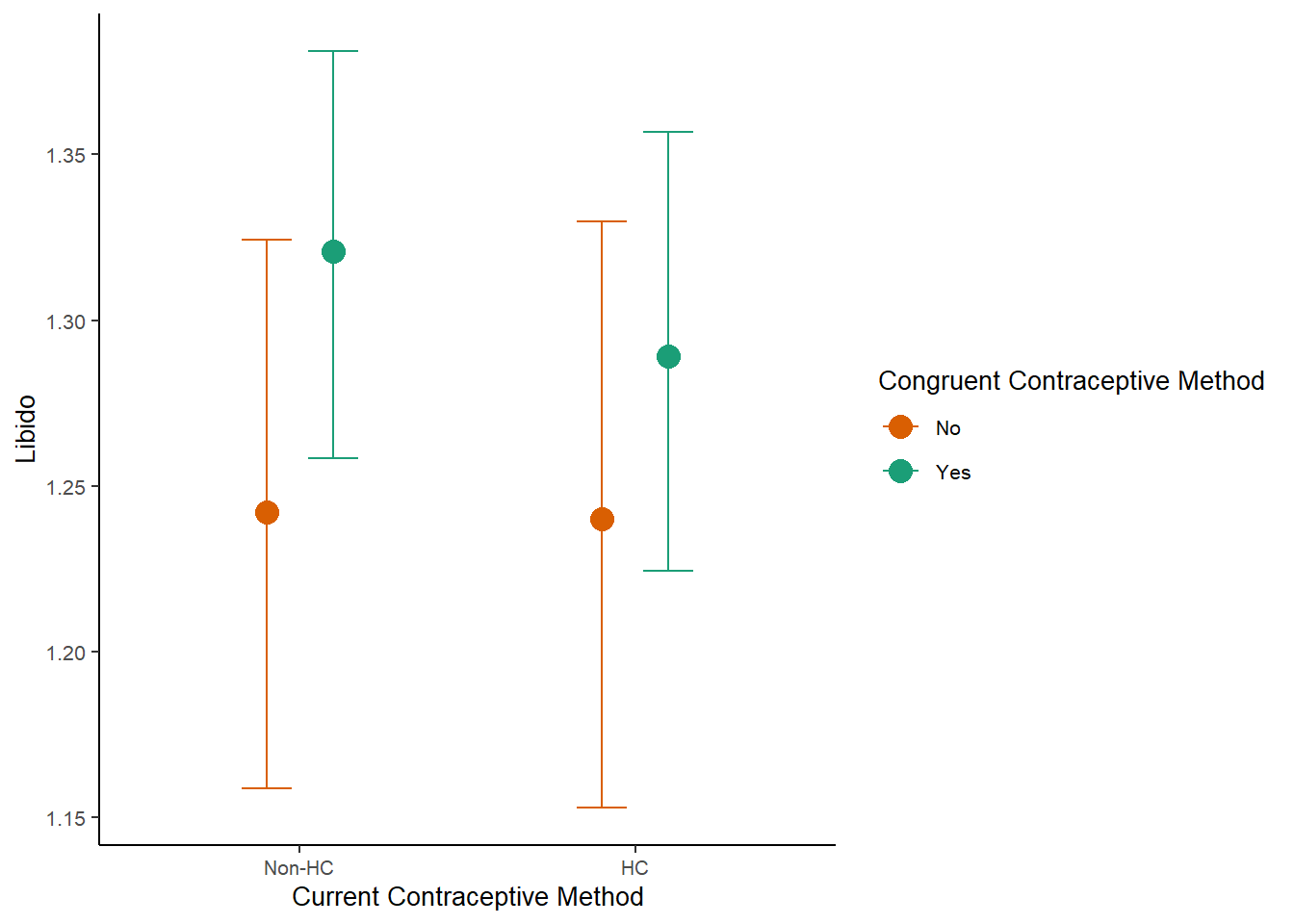
m_congruency_sexfreqpen = brm(diary_sex_active_sex_sum ~ offset(log(number_of_days)) +
contraception_hormonal * congruent_contraception,
data = data, family = poisson(),
file = "mrobust_congruency_sexfreqpen")
m_congruency_sexfreqpen_ce = m_congruency_sexfreqpen %>%
conditional_effects(., effects = "contraception_hormonal:congruent_contraception",
prob = 0.90,
conditions = data.frame(number_of_days = 1))
m_congruency_sexfreqpen_plot =
plot(m_congruency_sexfreqpen_ce, plot = FALSE)[[1]] +
labs(x = "Current Contraceptive Method",
y = "Frequency of Vaginal Intercourse\n(Probablity per Day)") +
scale_x_discrete(labels=c("Non-HC", "HC")) +
scale_color_manual(name = "Congruent Contraceptive Method", labels = c("No", "Yes"),
values=c("#D95F02", "#1B9E77")) +
scale_fill_manual(name = "Congruent Contraceptive Method", labels = c("No", "Yes"),
values=c("#D95F02", "#1B9E77")) +
apatheme
m_congruency_sexfreqpen_plot
m_congruency_masfreq = brm(diary_masturbation_sum ~ offset(log(number_of_days)) +
contraception_hormonal * congruent_contraception,
data = data, family = poisson(),
file = "mrobust_congruency_masfreq")
m_congruency_masfreq_ce = m_congruency_masfreq %>%
conditional_effects(., effects = "contraception_hormonal:congruent_contraception",
prob = 0.90,
conditions = data.frame(number_of_days = 1))
m_congruency_masfreq_plot =
plot(m_congruency_masfreq_ce, plot = FALSE)[[1]] +
labs(x = "Current Contraceptive Method",
y = "Frequency of Masturbation\n(Probablity per Day)") +
scale_x_discrete(labels=c("Non-HC", "HC")) +
scale_color_manual(name = "Congruent Contraceptive Method", labels = c("No", "Yes"),
values=c("#D95F02", "#1B9E77")) +
scale_fill_manual(name = "Congruent Contraceptive Method", labels = c("No", "Yes"),
values=c("#D95F02", "#1B9E77")) +
apatheme
m_congruency_masfreq_plot
Combine Plots
congruency_uncontrolled =
ggarrange(m_congruency_atrr_plot +
rremove("xlab"),
m_congruency_relsat_plot +
rremove("xlab"),
m_congruency_sexsat_plot +
rremove("xlab"),
m_congruency_libido_plot +
rremove("xlab"),
m_congruency_sexfreqpen_plot,
m_congruency_masfreq_plot,
common.legend = TRUE, legend = "bottom",
ncol = 2, nrow = 3,
align = "v")
congruency_uncontrolled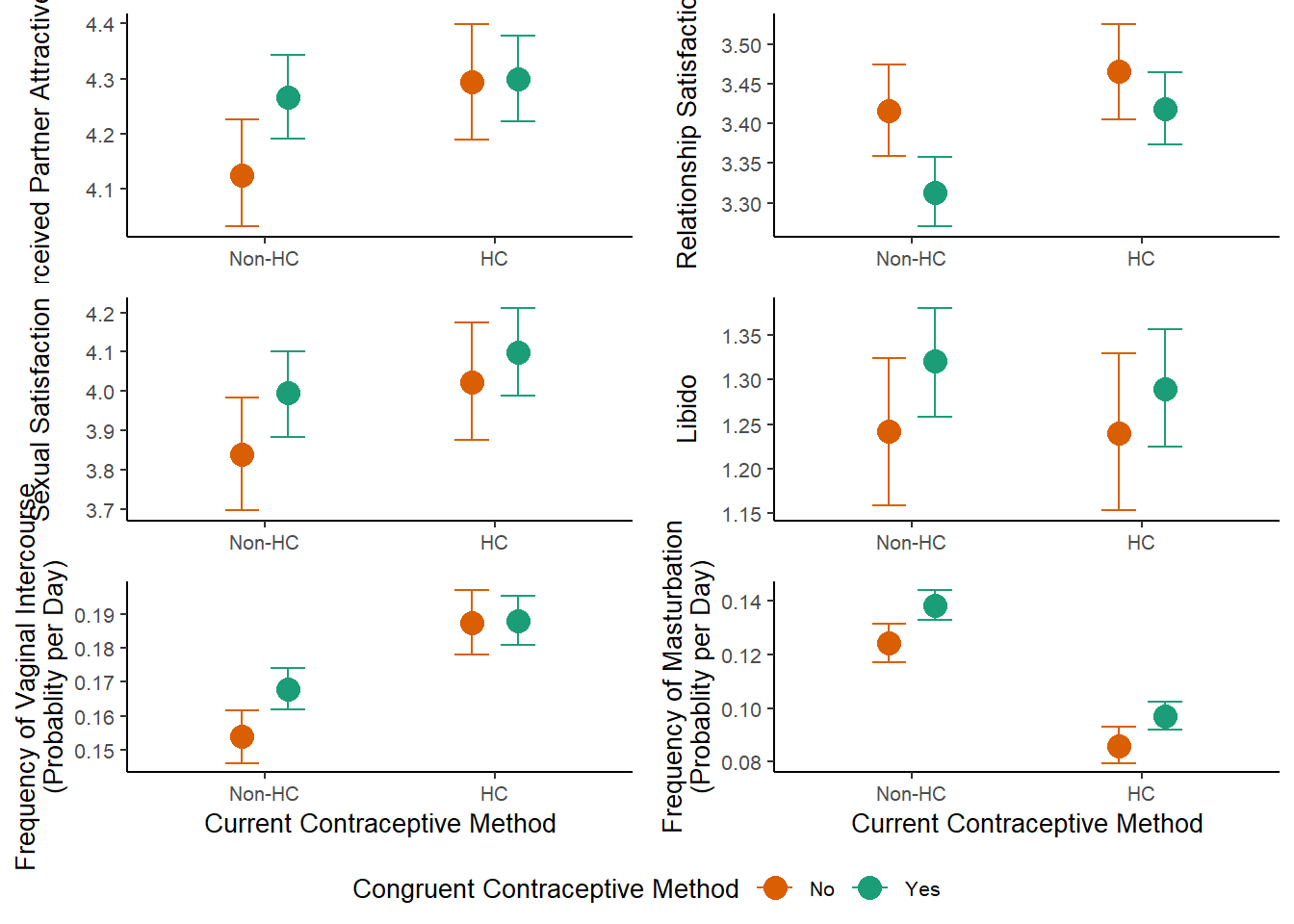
jpeg('Differences in current contraceptive method, congruent use of contraceptive method, and their interaction.jpg',
width = 800, height = 1000, quality = 100)
congruency_uncontrolled
dev.off()## png
## 2Controlled effects of (in)congruent use of hormonal contraceptives
Plots
m_congruency_atrr_controlled = brm(attractiveness_partner ~
contraception_hormonal * congruent_contraception +
age + net_income + relationship_duration_factor +
education_years +
bfi_extra + bfi_neuro + bfi_agree + bfi_consc + bfi_open +
religiosity,
data = data, family = gaussian(),
file = "m_congruency_atrr_controlled")
m_congruency_atrr_controlled_ce = m_congruency_atrr_controlled %>%
conditional_effects(., effects = "contraception_hormonal:congruent_contraception",
prob = 0.90)
m_congruency_atrr_controlled_plot =
plot(m_congruency_atrr_controlled_ce, plot = FALSE)[[1]] +
labs(x = "Current Contraceptive Method",
y = "Perceived Partner Attractiveness") +
scale_x_discrete(labels=c("Non-HC", "HC")) +
scale_color_manual(name = "Congruent Contraceptive Method", labels = c("No", "Yes"),
values=c("#D95F02", "#1B9E77")) +
scale_fill_manual(name = "Congruent Contraceptive Method", labels = c("No", "Yes"),
values=c("#D95F02", "#1B9E77")) +
apatheme
m_congruency_atrr_controlled_plot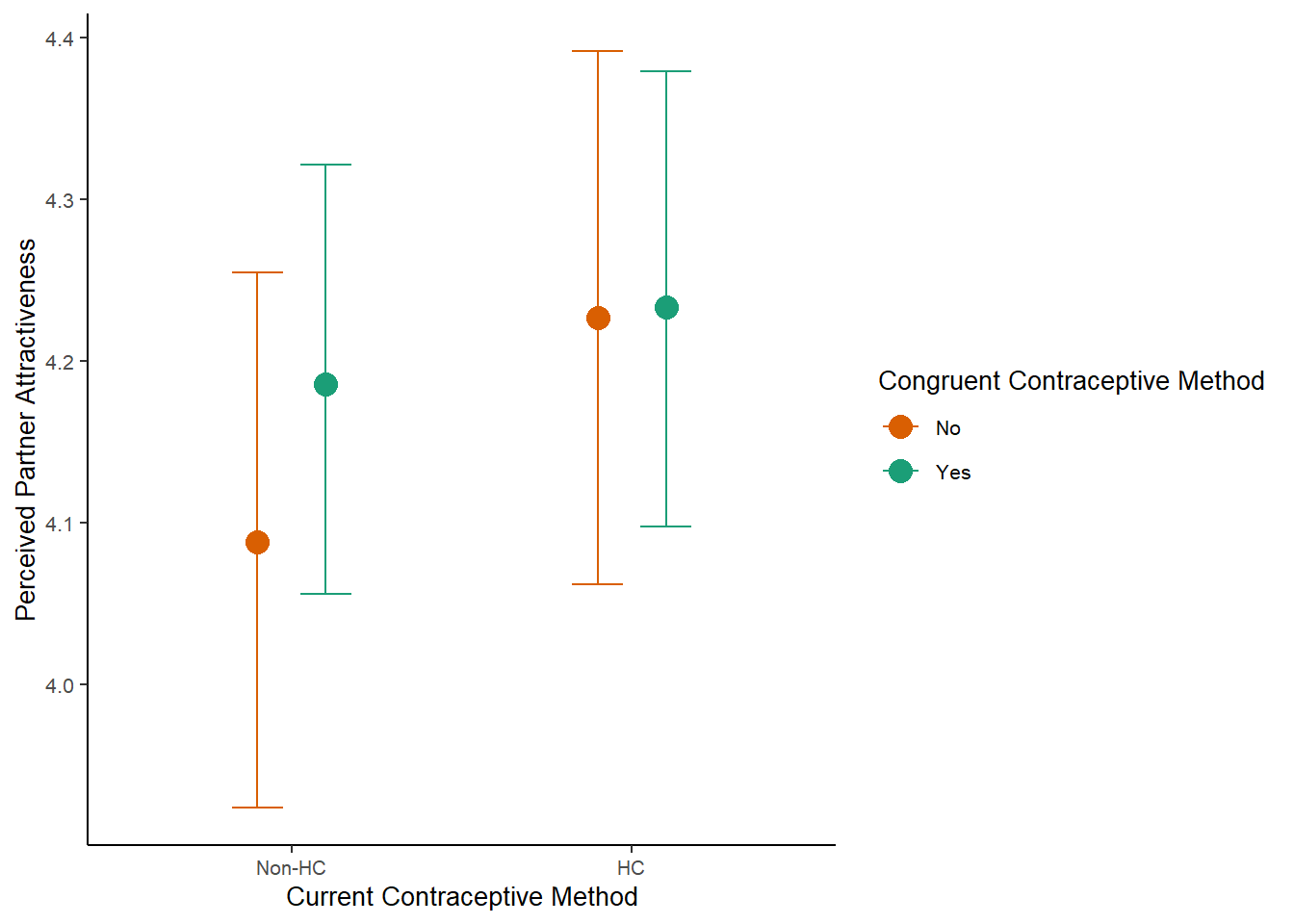
m_congruency_relsat_controlled = brm(relationship_satisfaction ~
contraception_hormonal * congruent_contraception +
age + net_income + relationship_duration_factor +
education_years +
bfi_extra + bfi_neuro + bfi_agree + bfi_consc + bfi_open +
religiosity,
data = data, family = gaussian(),
file = "m_congruency_relsat_controlled")
m_congruency_relsat_controlled_ce = m_congruency_relsat_controlled %>%
conditional_effects(., effects = "contraception_hormonal:congruent_contraception",
prob = 0.90)
m_congruency_relsat_controlled_plot =
plot(m_congruency_relsat_controlled_ce, plot = FALSE)[[1]] +
labs(x = "Current Contraceptive Method",
y = "Relationship Satisfaction") +
scale_x_discrete(labels=c("Non-HC", "HC")) +
scale_color_manual(name = "Congruent Contraceptive Method", labels = c("No", "Yes"),
values=c("#D95F02", "#1B9E77")) +
scale_fill_manual(name = "Congruent Contraceptive Method", labels = c("No", "Yes"),
values=c("#D95F02", "#1B9E77")) +
apatheme
m_congruency_relsat_controlled_plot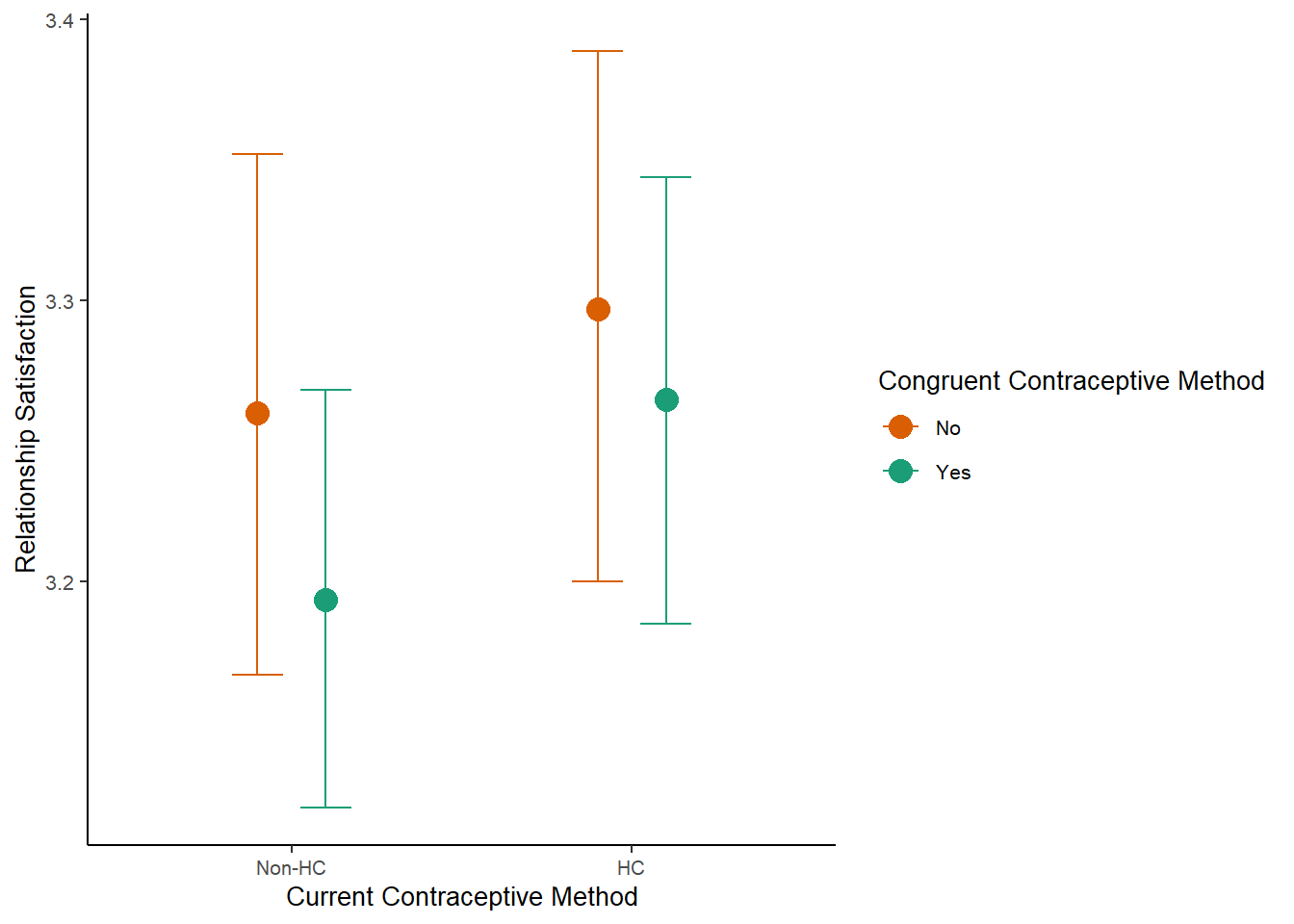
m_congruency_sexsat_controlled = brm(satisfaction_sexual_intercourse ~
contraception_hormonal * congruent_contraception +
age + net_income + relationship_duration_factor +
education_years +
bfi_extra + bfi_neuro + bfi_agree + bfi_consc + bfi_open +
religiosity,
data = data, family = gaussian(),
file = "m_congruency_sexsat_controlled")
m_congruency_sexsat_controlled_ce = m_congruency_sexsat_controlled %>%
conditional_effects(., effects = "contraception_hormonal:congruent_contraception",
prob = 0.90)
m_congruency_sexsat_controlled_plot =
plot(m_congruency_sexsat_controlled_ce, plot = FALSE)[[1]] +
labs(x = "Current Contraceptive Method",
y = "Sexual Satisfaction") +
scale_x_discrete(labels=c("Non-HC", "HC")) +
scale_color_manual(name = "Congruent Contraceptive Method", labels = c("No", "Yes"),
values=c("#D95F02", "#1B9E77")) +
scale_fill_manual(name = "Congruent Contraceptive Method", labels = c("No", "Yes"),
values=c("#D95F02", "#1B9E77")) +
apatheme
m_congruency_sexsat_controlled_plot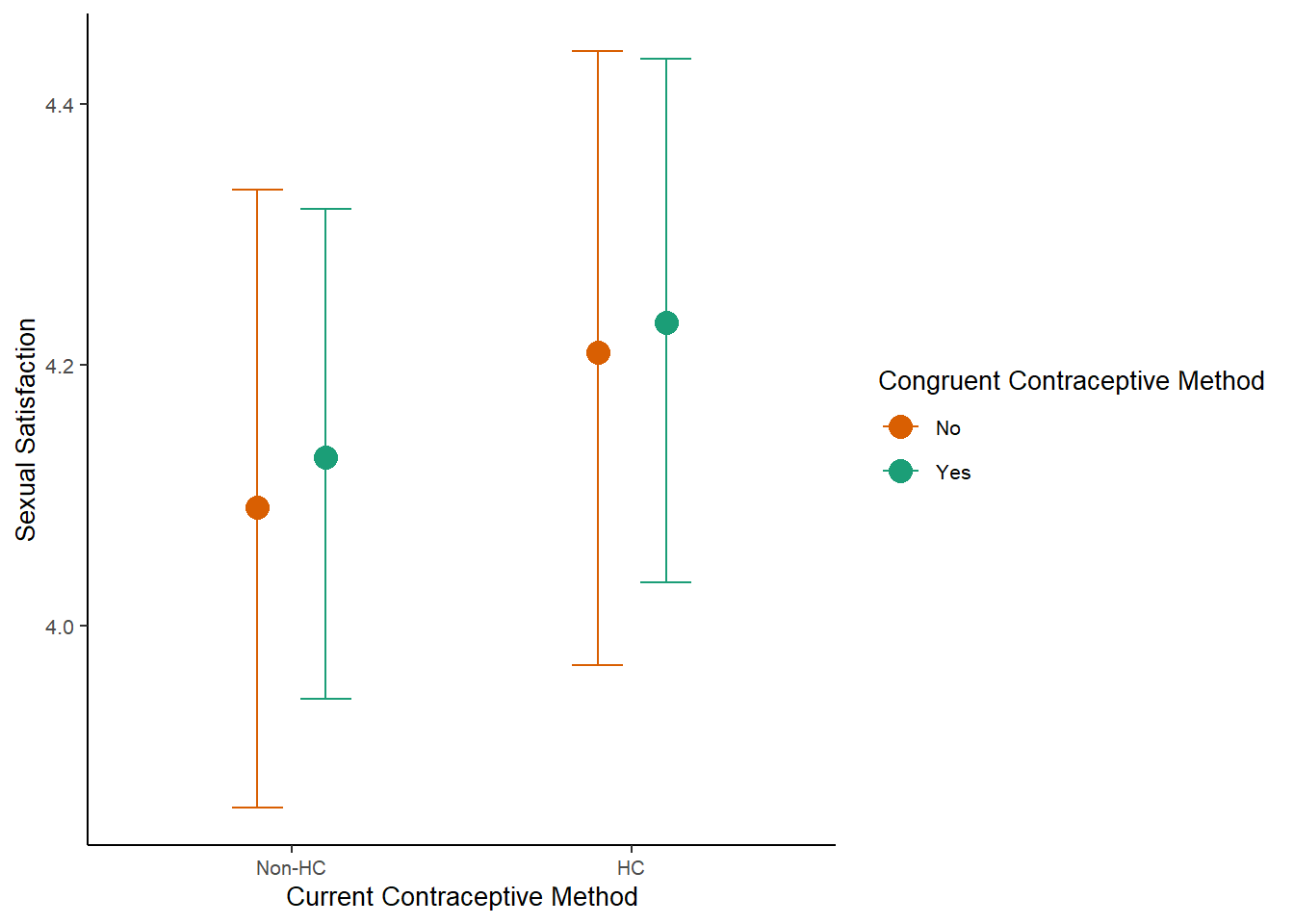
m_congruency_libido_controlled = brm(diary_libido_mean ~
contraception_hormonal * congruent_contraception +
age + net_income + relationship_duration_factor +
education_years +
bfi_extra + bfi_neuro + bfi_agree + bfi_consc + bfi_open +
religiosity,
data = data, family = gaussian(),
file = "m_congruency_libido_controlled")
m_congruency_libido_controlled_ce = m_congruency_libido_controlled %>%
conditional_effects(., effects = "contraception_hormonal:congruent_contraception",
prob = 0.90)
m_congruency_libido_controlled_plot =
plot(m_congruency_libido_controlled_ce, plot = FALSE)[[1]] +
labs(x = "Current Contraceptive Method",
y = "Libido") +
scale_x_discrete(labels=c("Non-HC", "HC")) +
scale_color_manual(name = "Congruent Contraceptive Method", labels = c("No", "Yes"),
values=c("#D95F02", "#1B9E77")) +
scale_fill_manual(name = "Congruent Contraceptive Method", labels = c("No", "Yes"),
values=c("#D95F02", "#1B9E77")) +
apatheme
m_congruency_libido_controlled_plot
m_congruency_sexfreqpen_controlled = brm(diary_sex_active_sex_sum ~
offset(log(number_of_days)) +
contraception_hormonal * congruent_contraception +
age + net_income + relationship_duration_factor +
education_years +
bfi_extra + bfi_neuro + bfi_agree + bfi_consc + bfi_open +
religiosity,
data = data, family = poisson(),
file = "mrobust_congruency_sexfreqpen_controlled")
m_congruency_sexfreqpen_controlled_ce = m_congruency_sexfreqpen_controlled %>%
conditional_effects(., effects = "contraception_hormonal:congruent_contraception",
prob = 0.90,
conditions = data.frame(number_of_days = 1))
m_congruency_sexfreqpen_controlled_plot =
plot(m_congruency_sexfreqpen_controlled_ce, plot = FALSE)[[1]] +
labs(x = "Current Contraceptive Method",
y = "Frequency of Vaginal Intercourse\n(Probability per Day)") +
scale_x_discrete(labels=c("Non-HC", "HC")) +
scale_color_manual(name = "Congruent Contraceptive Method", labels = c("No", "Yes"),
values=c("#D95F02", "#1B9E77")) +
scale_fill_manual(name = "Congruent Contraceptive Method", labels = c("No", "Yes"),
values=c("#D95F02", "#1B9E77")) +
apatheme
m_congruency_sexfreqpen_controlled_plot
m_congruency_masfreq_controlled = brm(diary_masturbation_sum ~
offset(log(number_of_days)) +
contraception_hormonal * congruent_contraception +
age + net_income + relationship_duration_factor +
education_years +
bfi_extra + bfi_neuro + bfi_agree + bfi_consc + bfi_open +
religiosity,
data = data, family = poisson(),
file = "mrobust_congruency_masfreq_controlled")
m_congruency_masfreq_controlled_ce = m_congruency_masfreq_controlled %>%
conditional_effects(., effects = "contraception_hormonal:congruent_contraception",
prob = 0.90,
conditions = data.frame(number_of_days = 1))
m_congruency_masfreq_controlled_plot =
plot(m_congruency_masfreq_controlled_ce, plot = FALSE)[[1]] +
labs(x = "Current Contraceptive Method",
y = "Frequency of Masturbation\n(Probability per Day)") +
scale_x_discrete(labels=c("Non-HC", "HC")) +
scale_color_manual(name = "Congruent Contraceptive Method", labels = c("No", "Yes"),
values=c("#D95F02", "#1B9E77")) +
scale_fill_manual(name = "Congruent Contraceptive Method", labels = c("No", "Yes"),
values=c("#D95F02", "#1B9E77")) +
apatheme
m_congruency_masfreq_controlled_plot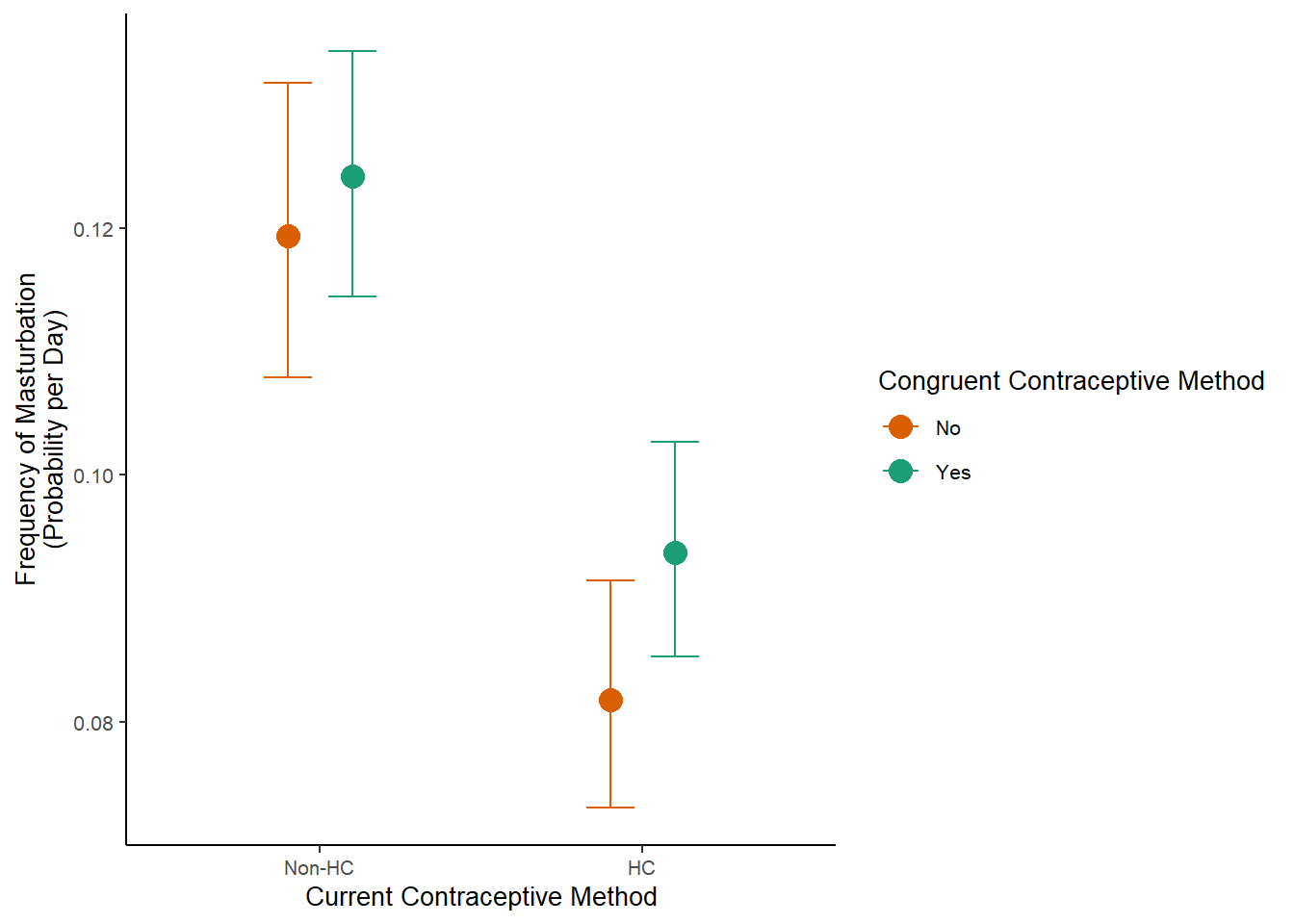
Combine Plots
congruency_controlled =
ggarrange(m_congruency_atrr_controlled_plot + rremove("xlab"),
m_congruency_relsat_controlled_plot + rremove("xlab"),
m_congruency_sexsat_controlled_plot + rremove("xlab"),
m_congruency_libido_controlled_plot + rremove("xlab"),
m_congruency_sexfreqpen_controlled_plot,
m_congruency_masfreq_controlled_plot,
common.legend = TRUE, legend = "bottom",
ncol = 2, nrow = 3,
align = "v")
congruency_controlled
jpeg('Differences in current contraceptive method, congruent use of contraceptive method, and their interaction in controlled models.jpg',
width = 800, height = 1000, quality = 100)
congruency_controlled
dev.off()## png
## 2LS0tDQp0aXRsZTogIlBsb3RzIEVmZmVjdHMgb2YgQ29udHJhY2VwdGlvbiINCm91dHB1dDoNCiAgaHRtbF9kb2N1bWVudDoNCiAgICB0b2M6IHRydWUNCiAgICB0b2NfZGVwdGg6IDQNCiAgICB0b2NfZmxvYXQ6IHRydWUNCiAgICBjb2RlX2ZvbGRpbmc6ICdoaWRlJw0KICAgIHNlbGZfY29udGFpbmVkOiBmYWxzZQ0KLS0tDQogIA0KICANCiMjIERhdGEgDQpgYGB7ciByZXN1bHRzPSdoaWRlJyxtZXNzYWdlPUYsd2FybmluZz1GfQ0Kc291cmNlKCIwX2hlbHBlcnMuUiIpDQoNCmxpYnJhcnkoZXh0cmFmb250KQ0KdGhlbWVfc2V0KHRoZW1lX3R1ZnRlKGJhc2Vfc2l6ZSA9IDEwLCBiYXNlX2ZhbWlseT0nQXJpYWwnKSkNCg0KI3Bsb3Qgc3BlY2lmaWNhdGlvbiBmb3IgcGFwZXI6DQojIGFwYXRoZW1lPXRoZW1lX2J3KCkrDQojICAgdGhlbWUocGFuZWwuZ3JpZC5tYWpvcj1lbGVtZW50X2JsYW5rKCksDQojICAgICAgICAgcGFuZWwuZ3JpZC5taW5vcj1lbGVtZW50X2JsYW5rKCksDQojICAgICAgICAgcGFuZWwuYm9yZGVyPWVsZW1lbnRfYmxhbmsoKSwNCiMgICAgICAgICBheGlzLmxpbmU9ZWxlbWVudF9saW5lKCksDQojICAgICAgICAgdGV4dCA9IGVsZW1lbnRfdGV4dChzaXplPTIwKSwNCiMgICAgICAgICBwbG90LnRpdGxlID0gZWxlbWVudF90ZXh0KHNpemUgPSAyMCkpDQoNCmxvYWQoImRhdGEvY2xlYW5lZF9zZWxlY3RlZF93cmFuZ2xlZC5yZGF0YSIpDQpgYGANCg0KDQojIyBTZWxlY3Rpb24gTW9kZWwgey50YWJzZXR9DQojIyMgSG9ybW9uYWwgQ29udHJhY2VwdGlvbiB7LnRhYnNldH0NCiMjIyMgU2ltcGxlIE1vZGVsDQpgYGB7cn0NCm1fc2VsZWN0aW9uX2hjX3NpbXBsZSA9IGJybShjb250cmFjZXB0aW9uX2hvcm1vbmFsIH4NCiAgICAgICAgICAgICAgICAgICAgICAgICAgICAgIGFnZSArIG5ldF9pbmNvbWUgKyByZWxhdGlvbnNoaXBfZHVyYXRpb25fZmFjdG9yLA0KICAgICAgICAgICAgICAgICAgICAgICAgICAgIGRhdGEgPSBkYXRhLA0KICAgICAgICAgICAgICAgICAgICAgICAgICAgIGZhbWlseSA9IGJlcm5vdWxsaSgicHJvYml0IiksDQogICAgICAgICAgICAgICAgICAgICAgICAgICAgZmlsZSA9ICJtX3NlbGVjdGlvbl9oY19zaW1wbGUiKQ0KDQptX3NlbGVjdGlvbl9oY19zaW1wbGVfcGxvdCA9IG1fc2VsZWN0aW9uX2hjX3NpbXBsZSAlPiUNCiAgc3ByZWFkX2RyYXdzKGJfYWdlLA0KICAgICAgICAgICAgICAgYl9uZXRfaW5jb21lZXVyb181MDBfMTAwMCwgYl9uZXRfaW5jb21lZXVyb18xMDAwXzIwMDAsIA0KICAgICAgICAgICAgICAgYl9uZXRfaW5jb21lZXVyb18yMDAwXzMwMDAsIGJfbmV0X2luY29tZWV1cm9fZ3RfMzAwMCwgYl9uZXRfaW5jb21lZG9udF90ZWxsLA0KICAgICAgICAgICAgICAgYl9yZWxhdGlvbnNoaXBfZHVyYXRpb25fZmFjdG9yUGFydG5lcmVkX3VwdG8xMm1vbnRocywNCiAgICAgICAgICAgICAgIGJfcmVsYXRpb25zaGlwX2R1cmF0aW9uX2ZhY3RvclBhcnRuZXJlZF91cHRvMjhtb250aHMsDQogICAgICAgICAgICAgICBiX3JlbGF0aW9uc2hpcF9kdXJhdGlvbl9mYWN0b3JQYXJ0bmVyZWRfdXB0bzUybW9udGhzLA0KICAgICAgICAgICAgICAgYl9yZWxhdGlvbnNoaXBfZHVyYXRpb25fZmFjdG9yUGFydG5lcmVkX21vcmV0aGFuNTJtb250aHMpICU+JQ0KICBwaXZvdF9sb25nZXIoY29scyA9IGMoYl9hZ2UsDQogICAgICAgICAgICAgICBiX25ldF9pbmNvbWVldXJvXzUwMF8xMDAwLCBiX25ldF9pbmNvbWVldXJvXzEwMDBfMjAwMCwgDQogICAgICAgICAgICAgICBiX25ldF9pbmNvbWVldXJvXzIwMDBfMzAwMCwgYl9uZXRfaW5jb21lZXVyb19ndF8zMDAwLCBiX25ldF9pbmNvbWVkb250X3RlbGwsDQogICAgICAgICAgICAgICBiX3JlbGF0aW9uc2hpcF9kdXJhdGlvbl9mYWN0b3JQYXJ0bmVyZWRfdXB0bzEybW9udGhzLA0KICAgICAgICAgICAgICAgYl9yZWxhdGlvbnNoaXBfZHVyYXRpb25fZmFjdG9yUGFydG5lcmVkX3VwdG8yOG1vbnRocywNCiAgICAgICAgICAgICAgIGJfcmVsYXRpb25zaGlwX2R1cmF0aW9uX2ZhY3RvclBhcnRuZXJlZF91cHRvNTJtb250aHMsDQogICAgICAgICAgICAgICBiX3JlbGF0aW9uc2hpcF9kdXJhdGlvbl9mYWN0b3JQYXJ0bmVyZWRfbW9yZXRoYW41Mm1vbnRocyksDQogICAgICAgICAgICAgICBuYW1lc190byA9ICJjb25kaXRpb24iLA0KICAgICAgICAgICAgICAgdmFsdWVzX3RvID0gInJfY29uZGl0aW9uIikgJT4lDQogIG11dGF0ZShjb25kaXRpb25fbWVhbiA9IHJfY29uZGl0aW9uLA0KICAgICAgICAgZ3JvdXAgPSBpZmVsc2UoY29uZGl0aW9uICVjb250YWlucyUgImJfcmVsYXRpb25zaGlwX2R1cmF0aW9uX2ZhY3RvciIsDQogICAgICAgICAgICAgICAgICAgICAgICAiUmVsYXRpb25zaGlwIER1cmF0aW9uIiwNCiAgICAgICAgICAgICAgICAgICAgICAgIGlmZWxzZShjb25kaXRpb24gJWNvbnRhaW5zJSAiYl9uZXRfaW5jb21lIiwNCiAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAiSW5jb21lIiwNCiAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICBOQSkpLA0KICAgICAgICAgY29uZGl0aW9uID0gaWZlbHNlKGNvbmRpdGlvbiA9PSAiYl9hZ2UiLCAiQWdlIiwNCiAgICAgICAgICAgICAgICBpZmVsc2UoY29uZGl0aW9uID09ICJiX25ldF9pbmNvbWVldXJvXzUwMF8xMDAwIiwgIjUwMCBcVTIwMTMgMSwwMDAgRXVybyBtb250aGx5IiwNCiAgICAgICAgICAgICAgICBpZmVsc2UoY29uZGl0aW9uID09ICJiX25ldF9pbmNvbWVldXJvXzEwMDBfMjAwMCIsICIxLDAwMCBcVTIwMTMgMiwwMDAgRXVybyBtb250aGx5IiwNCiAgICAgICAgICAgICAgICBpZmVsc2UoY29uZGl0aW9uID09ICJiX25ldF9pbmNvbWVldXJvXzIwMDBfMzAwMCIsICIyLDAwMCBcVTIwMTMgMywwMDAgRXVybyBtb250aGx5IiwNCiAgICAgICAgICAgICAgICBpZmVsc2UoY29uZGl0aW9uID09ICJiX25ldF9pbmNvbWVldXJvX2d0XzMwMDAiLCAiPjMsMDAwIEV1cm8gbW9udGhseSIsDQogICAgICAgICAgICAgICAgaWZlbHNlKGNvbmRpdGlvbiA9PSAiYl9uZXRfaW5jb21lZG9udF90ZWxsIiwgImRvIG5vdCBkaXNjbG9zZSIsDQogICAgICAgICAgICAgICAgaWZlbHNlKGNvbmRpdGlvbiA9PSAiYl9yZWxhdGlvbnNoaXBfZHVyYXRpb25fZmFjdG9yUGFydG5lcmVkX3VwdG8xMm1vbnRocyIsDQogICAgICAgICAgICAgICAgICAgICAgICIwIFxVMjAxMyAxMiBtb250aHMiLA0KICAgICAgICAgICAgICAgIGlmZWxzZShjb25kaXRpb24gPT0gImJfcmVsYXRpb25zaGlwX2R1cmF0aW9uX2ZhY3RvclBhcnRuZXJlZF91cHRvMjhtb250aHMiLA0KICAgICAgICAgICAgICAgICAgICAgICAiMTMgXFUyMDEzIDI4IG1vbnRocyIsDQogICAgICAgICAgICAgICAgaWZlbHNlKGNvbmRpdGlvbiA9PSAiYl9yZWxhdGlvbnNoaXBfZHVyYXRpb25fZmFjdG9yUGFydG5lcmVkX3VwdG81Mm1vbnRocyIsDQogICAgICAgICAgICAgICAgICAgICAgICIyOSBcVTIwMTMgNTIgbW9udGhzIiwNCiAgICAgICAgICAgICAgICBpZmVsc2UoY29uZGl0aW9uID09ICJiX3JlbGF0aW9uc2hpcF9kdXJhdGlvbl9mYWN0b3JQYXJ0bmVyZWRfbW9yZXRoYW41Mm1vbnRocyIsDQogICAgICAgICAgICAgICAgICAgICAgICI+NTIgbW9udGhzIiwNCiAgICAgICAgICAgICAgICAgICAgICAgY29uZGl0aW9uKSkpKSkpKSkpKSwNCiAgICAgICAgIGdyb3VwID0gaWZlbHNlKGlzLm5hKGdyb3VwKSwgY29uZGl0aW9uLCBncm91cCksDQogICAgICAgICBjb25kaXRpb24gPSBmYWN0b3IoY29uZGl0aW9uLCBsZXZlbHMgPSByZXYoYygiQWdlIiwNCiAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAiNTAwIFxVMjAxMyAxLDAwMCBFdXJvIG1vbnRobHkiLCAiMSwwMDAgXFUyMDEzIDIsMDAwIEV1cm8gbW9udGhseSIsDQogICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgIjIsMDAwIFxVMjAxMyAzLDAwMCBFdXJvIG1vbnRobHkiLCAiPjMsMDAwIEV1cm8gbW9udGhseSIsICJkbyBub3QgZGlzY2xvc2UiLA0KICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICIwIFxVMjAxMyAxMiBtb250aHMiLCAiMTMgXFUyMDEzIDI4IG1vbnRocyIsICIyOSBcVTIwMTMgNTIgbW9udGhzIiwNCiAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAiPjUyIG1vbnRocyIpKSkpICU+JQ0KICBnZ3Bsb3QoYWVzKHkgPSBjb25kaXRpb24sIHggPSBjb25kaXRpb25fbWVhbiwgY29sb3IgPSBncm91cCkpICsNCiAgc3RhdF9oYWxmZXllKC53aWR0aCA9IDAuOSkgKw0KICBnZW9tX3ZsaW5lKHhpbnRlcmNlcHQgPSAwLCBsaW5ldHlwZSA9ICJkb3R0ZWQiLCBzaXplID0gMSkgKw0KICBhcGF0aGVtZSArDQogIHRoZW1lKGxlZ2VuZC50aXRsZSA9IGVsZW1lbnRfYmxhbmsoKSkgKw0KICBsYWJzKHggPSAiRWZmZWN0IFNpemUgRXN0aW1hdGVzIiwgeSA9ICJQcmVkaWN0b3JzIikgKw0KICBzY2FsZV94X2NvbnRpbnVvdXMobGltaXRzID0gYygtMS4yLCAxLjkpLA0KICAgICAgICAgICAgICAgICAgICAgYnJlYWtzPSBjKC0xLjAsIC0wLjUsIDAsIDAuNSwgMS4wLCAxLjUpKSArDQogIHNjYWxlX2NvbG9yX21hbnVhbCh2YWx1ZXMgPSBjKCIjMUI5RTc3IiwgIiNEOTVGMDIiLCAiIzc1NzBCMyIpKSArDQogIGd1aWRlcyhjb2xvciA9IGd1aWRlX2xlZ2VuZChvdmVycmlkZS5hZXMgPSBsaXN0KHNpemUgPSAxMCkpKSArDQogIGdndGl0bGUoZXhwcmVzc2lvbihwYXN0ZShib2xkKCJTaW1wbGUgTW9kZWwgKCIpLA0KICAgICAgICAgICAgICAgICAgICAgICAgICAgYm9sZGl0YWxpYygibiIpLA0KICAgICAgICAgICAgICAgICAgICAgICAgICAgYm9sZCgiID0gMSwxNzkpIikpKSkgKw0KICB0aGVtZShwbG90LnRpdGxlID0gZWxlbWVudF90ZXh0KGhqdXN0ID0gLTAuMzIpKQ0KICAgICAgICAgIA0KbV9zZWxlY3Rpb25faGNfc2ltcGxlX3Bsb3QNCmBgYA0KDQojIyMjIENvbXBsZXggTW9kZWwNCmBgYHtyfQ0KbV9zZWxlY3Rpb25faGNfY29tcGxleCA9IGJybShjb250cmFjZXB0aW9uX2hvcm1vbmFsIH4NCiAgICAgICAgICAgICAgICAgICAgICAgICAgICAgIGFnZSArIG5ldF9pbmNvbWUgKyByZWxhdGlvbnNoaXBfZHVyYXRpb25fZmFjdG9yICsNCiAgICAgICAgICAgICAgICAgICAgICAgICAgICAgIGVkdWNhdGlvbl95ZWFycyArDQogICAgICAgICAgICAgICAgICAgICAgICAgICAgICBiZmlfZXh0cmEgKyBiZmlfbmV1cm8gKyBiZmlfYWdyZWUgKyBiZmlfY29uc2MgKyBiZmlfb3BlbiArDQogICAgICAgICAgICAgICAgICAgICAgICAgICAgICByZWxpZ2lvc2l0eSwNCiAgICAgICAgICAgICAgICAgICAgICAgICAgICBkYXRhID0gZGF0YSwNCiAgICAgICAgICAgICAgICAgICAgICAgICAgICBmYW1pbHkgPSBiZXJub3VsbGkoInByb2JpdCIpLA0KICAgICAgICAgICAgICAgICAgICAgICAgICAgIGZpbGUgPSAibV9zZWxlY3Rpb25faGNfY29tcGxleCIpDQoNCm1fc2VsZWN0aW9uX2hjX2NvbXBsZXhfcGxvdCA9IG1fc2VsZWN0aW9uX2hjX2NvbXBsZXggJT4lDQogIHNwcmVhZF9kcmF3cyhiX2FnZSwNCiAgICAgICAgICAgICAgIGJfbmV0X2luY29tZWV1cm9fNTAwXzEwMDAsIGJfbmV0X2luY29tZWV1cm9fMTAwMF8yMDAwLCANCiAgICAgICAgICAgICAgIGJfbmV0X2luY29tZWV1cm9fMjAwMF8zMDAwLCBiX25ldF9pbmNvbWVldXJvX2d0XzMwMDAsIGJfbmV0X2luY29tZWRvbnRfdGVsbCwNCiAgICAgICAgICAgICAgIGJfcmVsYXRpb25zaGlwX2R1cmF0aW9uX2ZhY3RvclBhcnRuZXJlZF91cHRvMTJtb250aHMsDQogICAgICAgICAgICAgICBiX3JlbGF0aW9uc2hpcF9kdXJhdGlvbl9mYWN0b3JQYXJ0bmVyZWRfdXB0bzI4bW9udGhzLA0KICAgICAgICAgICAgICAgYl9yZWxhdGlvbnNoaXBfZHVyYXRpb25fZmFjdG9yUGFydG5lcmVkX3VwdG81Mm1vbnRocywNCiAgICAgICAgICAgICAgIGJfcmVsYXRpb25zaGlwX2R1cmF0aW9uX2ZhY3RvclBhcnRuZXJlZF9tb3JldGhhbjUybW9udGhzLA0KICAgICAgICAgICAgICAgYl9lZHVjYXRpb25feWVhcnMsDQogICAgICAgICAgICAgICBiX2JmaV9leHRyYSwgYl9iZmlfbmV1cm8sIGJfYmZpX2FncmVlLCBiX2JmaV9jb25zYywgYl9iZmlfb3BlbiwgDQogICAgICAgICAgICAgICBiX3JlbGlnaW9zaXR5KSAlPiUNCiAgcGl2b3RfbG9uZ2VyKGNvbHMgPSBjKGJfYWdlLA0KICAgICAgICAgICAgICAgYl9uZXRfaW5jb21lZXVyb181MDBfMTAwMCwgYl9uZXRfaW5jb21lZXVyb18xMDAwXzIwMDAsIA0KICAgICAgICAgICAgICAgYl9uZXRfaW5jb21lZXVyb18yMDAwXzMwMDAsIGJfbmV0X2luY29tZWV1cm9fZ3RfMzAwMCwgYl9uZXRfaW5jb21lZG9udF90ZWxsLA0KICAgICAgICAgICAgICAgYl9yZWxhdGlvbnNoaXBfZHVyYXRpb25fZmFjdG9yUGFydG5lcmVkX3VwdG8xMm1vbnRocywNCiAgICAgICAgICAgICAgIGJfcmVsYXRpb25zaGlwX2R1cmF0aW9uX2ZhY3RvclBhcnRuZXJlZF91cHRvMjhtb250aHMsDQogICAgICAgICAgICAgICBiX3JlbGF0aW9uc2hpcF9kdXJhdGlvbl9mYWN0b3JQYXJ0bmVyZWRfdXB0bzUybW9udGhzLA0KICAgICAgICAgICAgICAgYl9yZWxhdGlvbnNoaXBfZHVyYXRpb25fZmFjdG9yUGFydG5lcmVkX21vcmV0aGFuNTJtb250aHMsDQogICAgICAgICAgICAgICBiX2VkdWNhdGlvbl95ZWFycywNCiAgICAgICAgICAgICAgIGJfYmZpX2V4dHJhLCBiX2JmaV9uZXVybywgYl9iZmlfYWdyZWUsIGJfYmZpX2NvbnNjLCBiX2JmaV9vcGVuLCANCiAgICAgICAgICAgICAgIGJfcmVsaWdpb3NpdHkpLA0KICAgICAgICAgICAgICAgbmFtZXNfdG8gPSAiY29uZGl0aW9uIiwNCiAgICAgICAgICAgICAgIHZhbHVlc190byA9ICJyX2NvbmRpdGlvbiIpICU+JQ0KICBtdXRhdGUoY29uZGl0aW9uX21lYW4gPSByX2NvbmRpdGlvbiwNCiAgICAgICAgIGdyb3VwID0gaWZlbHNlKGNvbmRpdGlvbiAlY29udGFpbnMlICJiX3JlbGF0aW9uc2hpcF9kdXJhdGlvbl9mYWN0b3IiLA0KICAgICAgICAgICAgICAgICAgICAgICAgIlJlbGF0aW9uc2hpcCBEdXJhdGlvbiIsDQogICAgICAgICAgICAgICAgICAgICAgICBpZmVsc2UoY29uZGl0aW9uICVjb250YWlucyUgImJfbmV0X2luY29tZSIsDQogICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgIkluY29tZSIsDQogICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgTkEpKSwNCiAgICAgICAgIGNvbmRpdGlvbiA9IGlmZWxzZShjb25kaXRpb24gPT0gImJfYWdlIiwgIkFnZSIsDQogICAgICAgICAgICAgICAgaWZlbHNlKGNvbmRpdGlvbiA9PSAiYl9uZXRfaW5jb21lZXVyb181MDBfMTAwMCIsICI1MDAgXFUyMDEzIDEsMDAwIEV1cm8gbW9udGhseSIsDQogICAgICAgICAgICAgICAgaWZlbHNlKGNvbmRpdGlvbiA9PSAiYl9uZXRfaW5jb21lZXVyb18xMDAwXzIwMDAiLCAiMSwwMDAgXFUyMDEzIDIsMDAwIEV1cm8gbW9udGhseSIsDQogICAgICAgICAgICAgICAgaWZlbHNlKGNvbmRpdGlvbiA9PSAiYl9uZXRfaW5jb21lZXVyb18yMDAwXzMwMDAiLCAiMiwwMDAgXFUyMDEzIDMsMDAwIEV1cm8gbW9udGhseSIsDQogICAgICAgICAgICAgICAgaWZlbHNlKGNvbmRpdGlvbiA9PSAiYl9uZXRfaW5jb21lZXVyb19ndF8zMDAwIiwgIj4zLDAwMCBFdXJvIG1vbnRobHkiLA0KICAgICAgICAgICAgICAgIGlmZWxzZShjb25kaXRpb24gPT0gImJfbmV0X2luY29tZWRvbnRfdGVsbCIsICJkbyBub3QgZGlzY2xvc2UiLA0KICAgICAgICAgICAgICAgIGlmZWxzZShjb25kaXRpb24gPT0gImJfcmVsYXRpb25zaGlwX2R1cmF0aW9uX2ZhY3RvclBhcnRuZXJlZF91cHRvMTJtb250aHMiLA0KICAgICAgICAgICAgICAgICAgICAgICAiMCBcVTIwMTMgMTIgbW9udGhzIiwNCiAgICAgICAgICAgICAgICBpZmVsc2UoY29uZGl0aW9uID09ICJiX3JlbGF0aW9uc2hpcF9kdXJhdGlvbl9mYWN0b3JQYXJ0bmVyZWRfdXB0bzI4bW9udGhzIiwNCiAgICAgICAgICAgICAgICAgICAgICAgIjEzIFxVMjAxMyAyOCBtb250aHMiLA0KICAgICAgICAgICAgICAgIGlmZWxzZShjb25kaXRpb24gPT0gImJfcmVsYXRpb25zaGlwX2R1cmF0aW9uX2ZhY3RvclBhcnRuZXJlZF91cHRvNTJtb250aHMiLA0KICAgICAgICAgICAgICAgICAgICAgICAiMjkgXFUyMDEzIDUyIG1vbnRocyIsDQogICAgICAgICAgICAgICAgaWZlbHNlKGNvbmRpdGlvbiA9PSAiYl9yZWxhdGlvbnNoaXBfZHVyYXRpb25fZmFjdG9yUGFydG5lcmVkX21vcmV0aGFuNTJtb250aHMiLA0KICAgICAgICAgICAgICAgICAgICAgICAiPjUyIG1vbnRocyIsDQogICAgICAgICAgICAgICAgaWZlbHNlKGNvbmRpdGlvbiA9PSAiYl9lZHVjYXRpb25feWVhcnMiLCAiWWVhcnMgb2YgRWR1Y2F0aW9uIiwNCiAgICAgICAgICAgICAgICBpZmVsc2UoY29uZGl0aW9uID09ICJiX2JmaV9leHRyYSIsICJFeHRyYXZlcnNpb24iLA0KICAgICAgICAgICAgICAgIGlmZWxzZShjb25kaXRpb24gPT0gImJfYmZpX25ldXJvIiwgIk5ldXJvdGljaXNtIiwNCiAgICAgICAgICAgICAgICBpZmVsc2UoY29uZGl0aW9uID09ICJiX2JmaV9hZ3JlZSIsICJBZ3JlZWFibGVuZXNzIiwNCiAgICAgICAgICAgICAgICBpZmVsc2UoY29uZGl0aW9uID09ICJiX2JmaV9jb25zYyIsICJDb25zY2llbnRpb3VzbmVzcyIsDQogICAgICAgICAgICAgICAgaWZlbHNlKGNvbmRpdGlvbiA9PSAiYl9iZmlfb3BlbiIsICJPcGVubmVzcyIsDQogICAgICAgICAgICAgICAgaWZlbHNlKGNvbmRpdGlvbiA9PSAiYl9yZWxpZ2lvc2l0eSIsICJSZWxpZ2lvc2l0eSIsDQogICAgICAgICAgICAgICAgICAgICAgIGNvbmRpdGlvbikpKSkpKSkpKSkpKSkpKSkpLA0KICAgICAgICAgZ3JvdXAgPSBpZmVsc2UoaXMubmEoZ3JvdXApLCBjb25kaXRpb24sIGdyb3VwKSwNCiAgICAgICAgIGNvbmRpdGlvbiA9IGZhY3Rvcihjb25kaXRpb24sIGxldmVscyA9IHJldihjKCJBZ2UiLA0KICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICI1MDAgXFUyMDEzIDEsMDAwIEV1cm8gbW9udGhseSIsICIxLDAwMCBcVTIwMTMgMiwwMDAgRXVybyBtb250aGx5IiwNCiAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAiMiwwMDAgXFUyMDEzIDMsMDAwIEV1cm8gbW9udGhseSIsICI+MywwMDAgRXVybyBtb250aGx5IiwgImRvIG5vdCBkaXNjbG9zZSIsDQogICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgIjAgXFUyMDEzIDEyIG1vbnRocyIsICIxMyBcVTIwMTMgMjggbW9udGhzIiwgIjI5IFxVMjAxMyA1MiBtb250aHMiLA0KICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICI+NTIgbW9udGhzIiwNCiAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAiWWVhcnMgb2YgRWR1Y2F0aW9uIiwNCiAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAiRXh0cmF2ZXJzaW9uIiwgIk5ldXJvdGljaXNtIiwgIkFncmVlYWJsZW5lc3MiLA0KICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAiQ29uc2NpZW50aW91c25lc3MiLCJPcGVubmVzcyIsIlJlbGlnaW9zaXR5IikpKSwNCiAgICAgICAgIGdyb3VwID0gZmFjdG9yKGdyb3VwLCBsZXZlbHMgPSBjKCJBZ2UiLCAiSW5jb21lIiwgIlJlbGF0aW9uc2hpcCBEdXJhdGlvbiIsDQogICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICANCiAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICJZZWFycyBvZiBFZHVjYXRpb24iLA0KICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICJFeHRyYXZlcnNpb24iLCAiTmV1cm90aWNpc20iLCAiQWdyZWVhYmxlbmVzcyIsDQogICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICJDb25zY2llbnRpb3VzbmVzcyIsIk9wZW5uZXNzIiwgIlJlbGlnaW9zaXR5IikpKSAlPiUNCiAgZ2dwbG90KGFlcyh5ID0gY29uZGl0aW9uLCB4ID0gY29uZGl0aW9uX21lYW4sIGNvbG9yID0gZ3JvdXApKSArDQogIHN0YXRfaGFsZmV5ZSgud2lkdGggPSAwLjkpICsNCiAgZ2VvbV92bGluZSh4aW50ZXJjZXB0ID0gMCwgbGluZXR5cGUgPSAiZG90dGVkIiwgc2l6ZSA9IDEpICsNCiAgYXBhdGhlbWUgKw0KICB0aGVtZShsZWdlbmQudGl0bGUgPSBlbGVtZW50X2JsYW5rKCkpICsNCiAgbGFicyh4ID0gIkVmZmVjdCBTaXplIEVzdGltYXRlcyIsIHkgPSAiUHJlZGljdG9ycyIpICsNCiAgc2NhbGVfeF9jb250aW51b3VzKGxpbWl0cyA9IGMoLTEuMiwgMS45KSwNCiAgICAgICAgICAgICAgICAgICAgIGJyZWFrcyA9IGMoLTEuMCwgLTAuNSwgMCwgMC41LCAxLjAsIDEuNSkpICsNCiAgc2NhbGVfY29sb3JfbWFudWFsKHZhbHVlcyA9IGMoIiMxQjlFNzciLCAiI0Q5NUYwMiIsICIjNzU3MEIzIiwgIiNFNzI5OEEiLCAiIzY2QTYxRSIsDQogICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICIjRTZBQjAyIiwgIiNBNjc2MUQiLCAiIzAwNzJCMiIsICIjQ0M3OUE3IiwgIiM1NkI0RTkiKSkgKw0KICBndWlkZXMoY29sb3IgPSBndWlkZV9sZWdlbmQob3ZlcnJpZGUuYWVzID0gbGlzdChzaXplID0gMTApKSkgKw0KICBnZ3RpdGxlKGV4cHJlc3Npb24ocGFzdGUoYm9sZCgiQ29tcGxleCBNb2RlbCAoIiksDQogICAgICAgICAgICAgICAgICAgICAgICAgICBib2xkaXRhbGljKCJuIiksDQogICAgICAgICAgICAgICAgICAgICAgICAgICBib2xkKCIgPSAxLDE3OSkiKSkpKSArDQogIHRoZW1lKHBsb3QudGl0bGUgPSBlbGVtZW50X3RleHQoaGp1c3QgPSAtMC4zMikpDQogICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICANCm1fc2VsZWN0aW9uX2hjX2NvbXBsZXhfcGxvdA0KYGBgDQoNCiMjIyMgQ29tYmluZSBQbG90cw0KYGBge3J9DQpzZWxlY3Rpb25faGMgPQ0KICBnZ2FycmFuZ2UobV9zZWxlY3Rpb25faGNfc2ltcGxlX3Bsb3QgKyB0aGVtZShsZWdlbmQucG9zaXRpb24gPSBjKDAuODUsMC41KSwNCiAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICBwbG90Lm1hcmdpbiA9IG1hcmdpbigxLDAsMCwwLCJjbSIpKSwNCiAgICAgICAgICBtX3NlbGVjdGlvbl9oY19jb21wbGV4X3Bsb3QgKyB0aGVtZShsZWdlbmQucG9zaXRpb24gPSBjKDAuODUsMC41KSwNCiAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICBwbG90Lm1hcmdpbiA9IG1hcmdpbigxLDAsMCwwLCJjbSIpKSwNCiAgICAgICAgICBoanVzdCA9IC0wLjEsDQogICAgICAgICAgbmNvbCA9IDEsIG5yb3cgPSAyLA0KICAgICAgICAgIGFsaWduID0gInYiKQ0KDQpzZWxlY3Rpb25faGMNCg0KanBlZygnRWZmZWN0IHNpemUgZXN0aW1hdGVzIGZvciBtb2RlbHMgd2l0aCBjdXJyZW50IGNvbnRyYWNlcHRpdmUgdXNlIGFzIGFuIG91dGNvbWUuanBnJywgDQogICAgIHdpZHRoID0gOTAwLCBoZWlnaHQgPSAxMDAwLCBxdWFsaXR5ID0gMTAwKQ0Kc2VsZWN0aW9uX2hjDQpkZXYub2ZmKCkNCmBgYA0KDQojIyMgKEluKWNvbmdydWVudCB1c2Ugb2YgaG9ybW9uYWwgY29udHJhY2VwdGl2ZXMgey50YWJzZXR9DQojIyMjIFNpbXBsZSBNb2RlbA0KYGBge3J9DQptX3NlbGVjdGlvbl9jb25ncnVlbnRfc2ltcGxlX29uY2VIQyA9IGJybShjb25ncnVlbnRfY29udHJhY2VwdGlvbiB+DQogICAgICAgICAgICAgICAgICAgICAgICAgICAgICBhZ2UgKyBuZXRfaW5jb21lICsgcmVsYXRpb25zaGlwX2R1cmF0aW9uX2ZhY3RvciwNCiAgICAgICAgICAgICAgICAgICAgICAgICAgICBkYXRhID0gZGF0YSAlPiUgZmlsdGVyKGNvbnRyYWNlcHRpb25fbWVldGluZ19wYXJ0bmVyID09ICJ5ZXMiKSwNCiAgICAgICAgICAgICAgICAgICAgICAgICAgICBmYW1pbHkgPSBiZXJub3VsbGkoInByb2JpdCIpLA0KICAgICAgICAgICAgICAgICAgICAgICAgICAgIGZpbGUgPSAibV9zZWxlY3Rpb25fY29uZ3J1ZW50X3NpbXBsZV9vbmNlSEMiKQ0KDQptX3NlbGVjdGlvbl9jb25ncnVlbnRfc2ltcGxlX29uY2Vub25IQyA9IGJybShjb25ncnVlbnRfY29udHJhY2VwdGlvbiB+DQogICAgICAgICAgICAgICAgICAgICAgICAgICAgICBhZ2UgKyBuZXRfaW5jb21lICsgcmVsYXRpb25zaGlwX2R1cmF0aW9uX2ZhY3RvciwNCiAgICAgICAgICAgICAgICAgICAgICAgICAgICBkYXRhID0gZGF0YSAlPiUgZmlsdGVyKGNvbnRyYWNlcHRpb25fbWVldGluZ19wYXJ0bmVyID09ICJubyIpLA0KICAgICAgICAgICAgICAgICAgICAgICAgICAgIGZhbWlseSA9IGJlcm5vdWxsaSgicHJvYml0IiksDQogICAgICAgICAgICAgICAgICAgICAgICAgICAgZmlsZSA9ICJtX3NlbGVjdGlvbl9jb25ncnVlbnRfc2ltcGxlX29uY2Vub25IQyIpDQoNCm1fc2VsZWN0aW9uX2NvbmdydWVudF9zaW1wbGVfb25jZUhDX3Bsb3QgPSBtX3NlbGVjdGlvbl9jb25ncnVlbnRfc2ltcGxlX29uY2VIQyAlPiUNCiAgc3ByZWFkX2RyYXdzKGJfYWdlLA0KICAgICAgICAgICAgICAgYl9uZXRfaW5jb21lZXVyb181MDBfMTAwMCwgYl9uZXRfaW5jb21lZXVyb18xMDAwXzIwMDAsIA0KICAgICAgICAgICAgICAgYl9uZXRfaW5jb21lZXVyb18yMDAwXzMwMDAsIGJfbmV0X2luY29tZWV1cm9fZ3RfMzAwMCwgYl9uZXRfaW5jb21lZG9udF90ZWxsLA0KICAgICAgICAgICAgICAgYl9yZWxhdGlvbnNoaXBfZHVyYXRpb25fZmFjdG9yUGFydG5lcmVkX3VwdG8yOG1vbnRocywNCiAgICAgICAgICAgICAgIGJfcmVsYXRpb25zaGlwX2R1cmF0aW9uX2ZhY3RvclBhcnRuZXJlZF91cHRvNTJtb250aHMsDQogICAgICAgICAgICAgICBiX3JlbGF0aW9uc2hpcF9kdXJhdGlvbl9mYWN0b3JQYXJ0bmVyZWRfbW9yZXRoYW41Mm1vbnRocykgJT4lDQogIHBpdm90X2xvbmdlcihjb2xzID0gYyhiX2FnZSwNCiAgICAgICAgICAgICAgIGJfbmV0X2luY29tZWV1cm9fNTAwXzEwMDAsIGJfbmV0X2luY29tZWV1cm9fMTAwMF8yMDAwLCANCiAgICAgICAgICAgICAgIGJfbmV0X2luY29tZWV1cm9fMjAwMF8zMDAwLCBiX25ldF9pbmNvbWVldXJvX2d0XzMwMDAsIGJfbmV0X2luY29tZWRvbnRfdGVsbCwNCiAgICAgICAgICAgICAgIGJfcmVsYXRpb25zaGlwX2R1cmF0aW9uX2ZhY3RvclBhcnRuZXJlZF91cHRvMjhtb250aHMsDQogICAgICAgICAgICAgICBiX3JlbGF0aW9uc2hpcF9kdXJhdGlvbl9mYWN0b3JQYXJ0bmVyZWRfdXB0bzUybW9udGhzLA0KICAgICAgICAgICAgICAgYl9yZWxhdGlvbnNoaXBfZHVyYXRpb25fZmFjdG9yUGFydG5lcmVkX21vcmV0aGFuNTJtb250aHMpLA0KICAgICAgICAgICAgICAgbmFtZXNfdG8gPSAiY29uZGl0aW9uIiwNCiAgICAgICAgICAgICAgIHZhbHVlc190byA9ICJyX2NvbmRpdGlvbiIpICU+JQ0KICBtdXRhdGUoY29uZGl0aW9uX21lYW4gPSByX2NvbmRpdGlvbiwNCiAgICAgICAgIGdyb3VwID0gaWZlbHNlKGNvbmRpdGlvbiAlY29udGFpbnMlICJiX3JlbGF0aW9uc2hpcF9kdXJhdGlvbl9mYWN0b3IiLA0KICAgICAgICAgICAgICAgICAgICAgICAgIlJlbGF0aW9uc2hpcCBEdXJhdGlvbiIsDQogICAgICAgICAgICAgICAgICAgICAgICBpZmVsc2UoY29uZGl0aW9uICVjb250YWlucyUgImJfbmV0X2luY29tZSIsDQogICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgIkluY29tZSIsDQogICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgTkEpKSwNCiAgICAgICAgIGNvbmRpdGlvbiA9IGlmZWxzZShjb25kaXRpb24gPT0gImJfYWdlIiwgIkFnZSIsDQogICAgICAgICAgICAgICAgaWZlbHNlKGNvbmRpdGlvbiA9PSAiYl9uZXRfaW5jb21lZXVyb181MDBfMTAwMCIsICI1MDAgXFUyMDEzIDEsMDAwIEV1cm8gbW9udGhseSIsDQogICAgICAgICAgICAgICAgaWZlbHNlKGNvbmRpdGlvbiA9PSAiYl9uZXRfaW5jb21lZXVyb18xMDAwXzIwMDAiLCAiMSwwMDAgXFUyMDEzIDIsMDAwIEV1cm8gbW9udGhseSIsDQogICAgICAgICAgICAgICAgaWZlbHNlKGNvbmRpdGlvbiA9PSAiYl9uZXRfaW5jb21lZXVyb18yMDAwXzMwMDAiLCAiMiwwMDAgXFUyMDEzIDMsMDAwIEV1cm8gbW9udGhseSIsDQogICAgICAgICAgICAgICAgaWZlbHNlKGNvbmRpdGlvbiA9PSAiYl9uZXRfaW5jb21lZXVyb19ndF8zMDAwIiwgIj4zLDAwMCBFdXJvIG1vbnRobHkiLA0KICAgICAgICAgICAgICAgIGlmZWxzZShjb25kaXRpb24gPT0gImJfbmV0X2luY29tZWRvbnRfdGVsbCIsICJkbyBub3QgZGlzY2xvc2UiLA0KICAgICAgICAgICAgICAgIGlmZWxzZShjb25kaXRpb24gPT0gImJfcmVsYXRpb25zaGlwX2R1cmF0aW9uX2ZhY3RvclBhcnRuZXJlZF91cHRvMTJtb250aHMiLA0KICAgICAgICAgICAgICAgICAgICAgICAiMCBcVTIwMTMgMTIgbW9udGhzIiwNCiAgICAgICAgICAgICAgICBpZmVsc2UoY29uZGl0aW9uID09ICJiX3JlbGF0aW9uc2hpcF9kdXJhdGlvbl9mYWN0b3JQYXJ0bmVyZWRfdXB0bzI4bW9udGhzIiwNCiAgICAgICAgICAgICAgICAgICAgICAgIjEzIFxVMjAxMyAyOCBtb250aHMiLA0KICAgICAgICAgICAgICAgIGlmZWxzZShjb25kaXRpb24gPT0gImJfcmVsYXRpb25zaGlwX2R1cmF0aW9uX2ZhY3RvclBhcnRuZXJlZF91cHRvNTJtb250aHMiLA0KICAgICAgICAgICAgICAgICAgICAgICAiMjkgXFUyMDEzIDUyIG1vbnRocyIsDQogICAgICAgICAgICAgICAgaWZlbHNlKGNvbmRpdGlvbiA9PSAiYl9yZWxhdGlvbnNoaXBfZHVyYXRpb25fZmFjdG9yUGFydG5lcmVkX21vcmV0aGFuNTJtb250aHMiLA0KICAgICAgICAgICAgICAgICAgICAgICAiPjUyIG1vbnRocyIsDQogICAgICAgICAgICAgICAgICAgICAgIGNvbmRpdGlvbikpKSkpKSkpKSksDQogICAgICAgICBncm91cCA9IGlmZWxzZShpcy5uYShncm91cCksIGNvbmRpdGlvbiwgZ3JvdXApLA0KICAgICAgICAgY29uZGl0aW9uID0gZmFjdG9yKGNvbmRpdGlvbiwgbGV2ZWxzID0gcmV2KGMoIkFnZSIsDQogICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgIjUwMCBcVTIwMTMgMSwwMDAgRXVybyBtb250aGx5IiwgIjEsMDAwIFxVMjAxMyAyLDAwMCBFdXJvIG1vbnRobHkiLA0KICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICIyLDAwMCBcVTIwMTMgMywwMDAgRXVybyBtb250aGx5IiwgIj4zLDAwMCBFdXJvIG1vbnRobHkiLCAiZG8gbm90IGRpc2Nsb3NlIiwNCiAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAiMTMgXFUyMDEzIDI4IG1vbnRocyIsICIyOSBcVTIwMTMgNTIgbW9udGhzIiwNCiAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAiPjUyIG1vbnRocyIpKSkpICU+JQ0KICBnZ3Bsb3QoYWVzKHkgPSBjb25kaXRpb24sIHggPSBjb25kaXRpb25fbWVhbiwgY29sb3IgPSBncm91cCkpICsNCiAgc3RhdF9oYWxmZXllKC53aWR0aCA9IC45MCkgKw0KICBnZW9tX3ZsaW5lKHhpbnRlcmNlcHQgPSAwLCBsaW5ldHlwZSA9ICJkb3R0ZWQiLCBzaXplID0gMSkgKw0KICBhcGF0aGVtZSArDQogIHRoZW1lKGxlZ2VuZC50aXRsZSA9IGVsZW1lbnRfYmxhbmsoKSkgKw0KICBsYWJzKHggPSAiRWZmZWN0IFNpemUgRXN0aW1hdGVzIiwgeSA9ICJQcmVkaWN0b3JzIikgKw0KICBzY2FsZV94X2NvbnRpbnVvdXMobGltaXRzID0gYygtMywgMyksDQogICAgICAgICAgICAgICAgICAgICBicmVha3M9IGMoLTM6MykpICsNCiAgc2NhbGVfY29sb3JfbWFudWFsKHZhbHVlcyA9IGMoIiMxQjlFNzciLCAiI0Q5NUYwMiIsICIjNzU3MEIzIikpICsNCiAgZ3VpZGVzKGNvbG9yID0gZ3VpZGVfbGVnZW5kKG92ZXJyaWRlLmFlcyA9IGxpc3Qoc2l6ZSA9IDUpKSkrDQogIGdndGl0bGUoZXhwcmVzc2lvbihwYXN0ZShib2xkKCJTaW1wbGUgTW9kZWwgSW5pdGlhbGx5IEhDIFVzZXIgKCIpLA0KICAgICAgICAgICAgICAgICAgICAgICAgICAgYm9sZGl0YWxpYygibiIpLA0KICAgICAgICAgICAgICAgICAgICAgICAgICAgYm9sZCgiID0gMzkwKSIpKSkpICsNCiAgdGhlbWUocGxvdC50aXRsZSA9IGVsZW1lbnRfdGV4dChoanVzdCA9IC0wLjM1KSkNCg0KbV9zZWxlY3Rpb25fY29uZ3J1ZW50X3NpbXBsZV9vbmNlSENfcGxvdA0KDQoNCm1fc2VsZWN0aW9uX2NvbmdydWVudF9zaW1wbGVfb25jZW5vbkhDX3Bsb3QgPSBtX3NlbGVjdGlvbl9jb25ncnVlbnRfc2ltcGxlX29uY2Vub25IQyAlPiUNCiAgc3ByZWFkX2RyYXdzKGJfYWdlLA0KICAgICAgICAgICAgICAgYl9uZXRfaW5jb21lZXVyb181MDBfMTAwMCwgYl9uZXRfaW5jb21lZXVyb18xMDAwXzIwMDAsIA0KICAgICAgICAgICAgICAgYl9uZXRfaW5jb21lZXVyb18yMDAwXzMwMDAsIGJfbmV0X2luY29tZWV1cm9fZ3RfMzAwMCwgYl9uZXRfaW5jb21lZG9udF90ZWxsLA0KICAgICAgICAgICAgICAgYl9yZWxhdGlvbnNoaXBfZHVyYXRpb25fZmFjdG9yUGFydG5lcmVkX3VwdG8yOG1vbnRocywNCiAgICAgICAgICAgICAgIGJfcmVsYXRpb25zaGlwX2R1cmF0aW9uX2ZhY3RvclBhcnRuZXJlZF91cHRvNTJtb250aHMsDQogICAgICAgICAgICAgICBiX3JlbGF0aW9uc2hpcF9kdXJhdGlvbl9mYWN0b3JQYXJ0bmVyZWRfbW9yZXRoYW41Mm1vbnRocykgJT4lDQogIHBpdm90X2xvbmdlcihjb2xzID0gYyhiX2FnZSwNCiAgICAgICAgICAgICAgIGJfbmV0X2luY29tZWV1cm9fNTAwXzEwMDAsIGJfbmV0X2luY29tZWV1cm9fMTAwMF8yMDAwLCANCiAgICAgICAgICAgICAgIGJfbmV0X2luY29tZWV1cm9fMjAwMF8zMDAwLCBiX25ldF9pbmNvbWVldXJvX2d0XzMwMDAsIGJfbmV0X2luY29tZWRvbnRfdGVsbCwNCiAgICAgICAgICAgICAgIGJfcmVsYXRpb25zaGlwX2R1cmF0aW9uX2ZhY3RvclBhcnRuZXJlZF91cHRvMjhtb250aHMsDQogICAgICAgICAgICAgICBiX3JlbGF0aW9uc2hpcF9kdXJhdGlvbl9mYWN0b3JQYXJ0bmVyZWRfdXB0bzUybW9udGhzLA0KICAgICAgICAgICAgICAgYl9yZWxhdGlvbnNoaXBfZHVyYXRpb25fZmFjdG9yUGFydG5lcmVkX21vcmV0aGFuNTJtb250aHMpLA0KICAgICAgICAgICAgICAgbmFtZXNfdG8gPSAiY29uZGl0aW9uIiwNCiAgICAgICAgICAgICAgIHZhbHVlc190byA9ICJyX2NvbmRpdGlvbiIpICU+JQ0KICBtdXRhdGUoY29uZGl0aW9uX21lYW4gPSByX2NvbmRpdGlvbiwNCiAgICAgICAgIGdyb3VwID0gaWZlbHNlKGNvbmRpdGlvbiAlY29udGFpbnMlICJiX3JlbGF0aW9uc2hpcF9kdXJhdGlvbl9mYWN0b3IiLA0KICAgICAgICAgICAgICAgICAgICAgICAgIlJlbGF0aW9uc2hpcCBEdXJhdGlvbiIsDQogICAgICAgICAgICAgICAgICAgICAgICBpZmVsc2UoY29uZGl0aW9uICVjb250YWlucyUgImJfbmV0X2luY29tZSIsDQogICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgIkluY29tZSIsDQogICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgTkEpKSwNCiAgICAgICAgIGNvbmRpdGlvbiA9IGlmZWxzZShjb25kaXRpb24gPT0gImJfYWdlIiwgIkFnZSIsDQogICAgICAgICAgICAgICAgaWZlbHNlKGNvbmRpdGlvbiA9PSAiYl9uZXRfaW5jb21lZXVyb181MDBfMTAwMCIsICI1MDAgXFUyMDEzIDEsMDAwIEV1cm8gbW9udGhseSIsDQogICAgICAgICAgICAgICAgaWZlbHNlKGNvbmRpdGlvbiA9PSAiYl9uZXRfaW5jb21lZXVyb18xMDAwXzIwMDAiLCAiMSwwMDAgXFUyMDEzIDIsMDAwIEV1cm8gbW9udGhseSIsDQogICAgICAgICAgICAgICAgaWZlbHNlKGNvbmRpdGlvbiA9PSAiYl9uZXRfaW5jb21lZXVyb18yMDAwXzMwMDAiLCAiMiwwMDAgXFUyMDEzIDMsMDAwIEV1cm8gbW9udGhseSIsDQogICAgICAgICAgICAgICAgaWZlbHNlKGNvbmRpdGlvbiA9PSAiYl9uZXRfaW5jb21lZXVyb19ndF8zMDAwIiwgIj4zLDAwMCBFdXJvIG1vbnRobHkiLA0KICAgICAgICAgICAgICAgIGlmZWxzZShjb25kaXRpb24gPT0gImJfbmV0X2luY29tZWRvbnRfdGVsbCIsICJkbyBub3QgZGlzY2xvc2UiLA0KICAgICAgICAgICAgICAgIGlmZWxzZShjb25kaXRpb24gPT0gImJfcmVsYXRpb25zaGlwX2R1cmF0aW9uX2ZhY3RvclBhcnRuZXJlZF91cHRvMTJtb250aHMiLA0KICAgICAgICAgICAgICAgICAgICAgICAiMCBcVTIwMTMgMTIgbW9udGhzIiwNCiAgICAgICAgICAgICAgICBpZmVsc2UoY29uZGl0aW9uID09ICJiX3JlbGF0aW9uc2hpcF9kdXJhdGlvbl9mYWN0b3JQYXJ0bmVyZWRfdXB0bzI4bW9udGhzIiwNCiAgICAgICAgICAgICAgICAgICAgICAgIjEzIFxVMjAxMyAyOCBtb250aHMiLA0KICAgICAgICAgICAgICAgIGlmZWxzZShjb25kaXRpb24gPT0gImJfcmVsYXRpb25zaGlwX2R1cmF0aW9uX2ZhY3RvclBhcnRuZXJlZF91cHRvNTJtb250aHMiLA0KICAgICAgICAgICAgICAgICAgICAgICAiMjkgXFUyMDEzIDUyIG1vbnRocyIsDQogICAgICAgICAgICAgICAgaWZlbHNlKGNvbmRpdGlvbiA9PSAiYl9yZWxhdGlvbnNoaXBfZHVyYXRpb25fZmFjdG9yUGFydG5lcmVkX21vcmV0aGFuNTJtb250aHMiLA0KICAgICAgICAgICAgICAgICAgICAgICAiPjUyIG1vbnRocyIsDQogICAgICAgICAgICAgICAgICAgICAgIGNvbmRpdGlvbikpKSkpKSkpKSksDQogICAgICAgICBncm91cCA9IGlmZWxzZShpcy5uYShncm91cCksIGNvbmRpdGlvbiwgZ3JvdXApLA0KICAgICAgICAgY29uZGl0aW9uID0gZmFjdG9yKGNvbmRpdGlvbiwgbGV2ZWxzID0gcmV2KGMoIkFnZSIsDQogICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgIjUwMCBcVTIwMTMgMSwwMDAgRXVybyBtb250aGx5IiwgIjEsMDAwIFxVMjAxMyAyLDAwMCBFdXJvIG1vbnRobHkiLA0KICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICIyLDAwMCBcVTIwMTMgMywwMDAgRXVybyBtb250aGx5IiwgIj4zLDAwMCBFdXJvIG1vbnRobHkiLCAiZG8gbm90IGRpc2Nsb3NlIiwNCiAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAiMTMgXFUyMDEzIDI4IG1vbnRocyIsICIyOSBcVTIwMTMgNTIgbW9udGhzIiwNCiAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAiPjUyIG1vbnRocyIpKSkpICU+JQ0KICBnZ3Bsb3QoYWVzKHkgPSBjb25kaXRpb24sIHggPSBjb25kaXRpb25fbWVhbiwgY29sb3IgPSBncm91cCkpICsNCiAgc3RhdF9oYWxmZXllKC53aWR0aCA9IC45MCkgKw0KICBnZW9tX3ZsaW5lKHhpbnRlcmNlcHQgPSAwLCBsaW5ldHlwZSA9ICJkb3R0ZWQiLCBzaXplID0gMSkgKw0KICBhcGF0aGVtZSArDQogIHRoZW1lKGxlZ2VuZC50aXRsZSA9IGVsZW1lbnRfYmxhbmsoKSkgKw0KICBsYWJzKHggPSAiRWZmZWN0IFNpemUgRXN0aW1hdGVzIiwgeSA9ICJQcmVkaWN0b3JzIikgKw0KICBzY2FsZV94X2NvbnRpbnVvdXMobGltaXRzID0gYygtMywgMyksDQogICAgICAgICAgICAgICAgICAgICBicmVha3M9IGMoLTM6MykpICsNCiAgc2NhbGVfY29sb3JfbWFudWFsKHZhbHVlcyA9IGMoIiMxQjlFNzciLCAiI0Q5NUYwMiIsICIjNzU3MEIzIikpICsNCiAgZ3VpZGVzKGNvbG9yID0gZ3VpZGVfbGVnZW5kKG92ZXJyaWRlLmFlcyA9IGxpc3Qoc2l6ZSA9IDUpKSkgKw0KICBnZ3RpdGxlKGV4cHJlc3Npb24ocGFzdGUoYm9sZCgiU2ltcGxlIE1vZGVsIEluaXRpYWxseSBOb24tSEMgVXNlciAoIiksDQogICAgICAgICAgICAgICAgICAgICAgICAgICBib2xkaXRhbGljKCJuIiksDQogICAgICAgICAgICAgICAgICAgICAgICAgICBib2xkKCIgPSAzODQpIikpKSkgKw0KICB0aGVtZShwbG90LnRpdGxlID0gZWxlbWVudF90ZXh0KGhqdXN0ID0gLTAuMzUpKQ0KbV9zZWxlY3Rpb25fY29uZ3J1ZW50X3NpbXBsZV9vbmNlbm9uSENfcGxvdA0KYGBgDQoNCiMjIyMgQ29tcGxleCBNb2RlbA0KYGBge3J9DQptX3NlbGVjdGlvbl9jb25ncnVlbnRfY29tcGxleF9vbmNlSEMgPSBicm0oY29uZ3J1ZW50X2NvbnRyYWNlcHRpb24gfg0KICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgYWdlICsgbmV0X2luY29tZSArIHJlbGF0aW9uc2hpcF9kdXJhdGlvbl9mYWN0b3IgKw0KICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgZWR1Y2F0aW9uX3llYXJzICsNCiAgICAgICAgICAgICAgICAgICAgICAgICAgICAgIGJmaV9leHRyYSArIGJmaV9uZXVybyArIGJmaV9hZ3JlZSArIGJmaV9jb25zYyArIGJmaV9vcGVuICsNCiAgICAgICAgICAgICAgICAgICAgICAgICAgICAgIHJlbGlnaW9zaXR5LA0KICAgICAgICAgICAgICAgICAgICAgICAgICAgIGRhdGEgPSBkYXRhICU+JSBmaWx0ZXIoY29udHJhY2VwdGlvbl9tZWV0aW5nX3BhcnRuZXIgPT0gInllcyIpLA0KICAgICAgICAgICAgICAgICAgICAgICAgICAgIGZhbWlseSA9IGJlcm5vdWxsaSgicHJvYml0IiksDQogICAgICAgICAgICAgICAgICAgICAgICAgICAgZmlsZSA9ICJtX3NlbGVjdGlvbl9jb25ncnVlbnRfY29tcGxleF9vbmNlSEMiKQ0KDQptX3NlbGVjdGlvbl9jb25ncnVlbnRfY29tcGxleF9vbmNlbm9uSEMgPSBicm0oY29uZ3J1ZW50X2NvbnRyYWNlcHRpb24gfg0KICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgYWdlICsgbmV0X2luY29tZSArIHJlbGF0aW9uc2hpcF9kdXJhdGlvbl9mYWN0b3IgKw0KICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgZWR1Y2F0aW9uX3llYXJzICsNCiAgICAgICAgICAgICAgICAgICAgICAgICAgICAgIGJmaV9leHRyYSArIGJmaV9uZXVybyArIGJmaV9hZ3JlZSArIGJmaV9jb25zYyArIGJmaV9vcGVuICsNCiAgICAgICAgICAgICAgICAgICAgICAgICAgICAgIHJlbGlnaW9zaXR5LA0KICAgICAgICAgICAgICAgICAgICAgICAgICAgIGRhdGEgPSBkYXRhICU+JSBmaWx0ZXIoY29udHJhY2VwdGlvbl9tZWV0aW5nX3BhcnRuZXIgPT0gIm5vIiksDQogICAgICAgICAgICAgICAgICAgICAgICAgICAgZmFtaWx5ID0gYmVybm91bGxpKCJwcm9iaXQiKSwNCiAgICAgICAgICAgICAgICAgICAgICAgICAgICBmaWxlID0gIm1fc2VsZWN0aW9uX2NvbmdydWVudF9jb21wbGV4X29uY2Vub25IQyIpDQoNCm1fc2VsZWN0aW9uX2NvbmdydWVudF9jb21wbGV4X29uY2VIQ19wbG90ID0gbV9zZWxlY3Rpb25fY29uZ3J1ZW50X2NvbXBsZXhfb25jZUhDICU+JQ0KICBzcHJlYWRfZHJhd3MoYl9hZ2UsDQogICAgICAgICAgICAgICBiX25ldF9pbmNvbWVldXJvXzUwMF8xMDAwLCBiX25ldF9pbmNvbWVldXJvXzEwMDBfMjAwMCwgDQogICAgICAgICAgICAgICBiX25ldF9pbmNvbWVldXJvXzIwMDBfMzAwMCwgYl9uZXRfaW5jb21lZXVyb19ndF8zMDAwLCBiX25ldF9pbmNvbWVkb250X3RlbGwsDQogICAgICAgICAgICAgICBiX3JlbGF0aW9uc2hpcF9kdXJhdGlvbl9mYWN0b3JQYXJ0bmVyZWRfdXB0bzI4bW9udGhzLA0KICAgICAgICAgICAgICAgYl9yZWxhdGlvbnNoaXBfZHVyYXRpb25fZmFjdG9yUGFydG5lcmVkX3VwdG81Mm1vbnRocywNCiAgICAgICAgICAgICAgIGJfcmVsYXRpb25zaGlwX2R1cmF0aW9uX2ZhY3RvclBhcnRuZXJlZF9tb3JldGhhbjUybW9udGhzLA0KICAgICAgICAgICAgICAgYl9lZHVjYXRpb25feWVhcnMsDQogICAgICAgICAgICAgICBiX2JmaV9leHRyYSwgYl9iZmlfbmV1cm8sIGJfYmZpX2FncmVlLCBiX2JmaV9jb25zYywgYl9iZmlfb3BlbiwgDQogICAgICAgICAgICAgICBiX3JlbGlnaW9zaXR5KSAlPiUNCiAgcGl2b3RfbG9uZ2VyKGNvbHMgPSBjKGJfYWdlLA0KICAgICAgICAgICAgICAgYl9uZXRfaW5jb21lZXVyb181MDBfMTAwMCwgYl9uZXRfaW5jb21lZXVyb18xMDAwXzIwMDAsIA0KICAgICAgICAgICAgICAgYl9uZXRfaW5jb21lZXVyb18yMDAwXzMwMDAsIGJfbmV0X2luY29tZWV1cm9fZ3RfMzAwMCwgYl9uZXRfaW5jb21lZG9udF90ZWxsLA0KICAgICAgICAgICAgICAgYl9yZWxhdGlvbnNoaXBfZHVyYXRpb25fZmFjdG9yUGFydG5lcmVkX3VwdG8yOG1vbnRocywNCiAgICAgICAgICAgICAgIGJfcmVsYXRpb25zaGlwX2R1cmF0aW9uX2ZhY3RvclBhcnRuZXJlZF91cHRvNTJtb250aHMsDQogICAgICAgICAgICAgICBiX3JlbGF0aW9uc2hpcF9kdXJhdGlvbl9mYWN0b3JQYXJ0bmVyZWRfbW9yZXRoYW41Mm1vbnRocywNCiAgICAgICAgICAgICAgIGJfZWR1Y2F0aW9uX3llYXJzLA0KICAgICAgICAgICAgICAgYl9iZmlfZXh0cmEsIGJfYmZpX25ldXJvLCBiX2JmaV9hZ3JlZSwgYl9iZmlfY29uc2MsIGJfYmZpX29wZW4sIA0KICAgICAgICAgICAgICAgYl9yZWxpZ2lvc2l0eSksDQogICAgICAgICAgICAgICBuYW1lc190byA9ICJjb25kaXRpb24iLA0KICAgICAgICAgICAgICAgdmFsdWVzX3RvID0gInJfY29uZGl0aW9uIikgJT4lDQogIG11dGF0ZShjb25kaXRpb25fbWVhbiA9IHJfY29uZGl0aW9uLA0KICAgICAgICAgZ3JvdXAgPSBpZmVsc2UoY29uZGl0aW9uICVjb250YWlucyUgImJfcmVsYXRpb25zaGlwX2R1cmF0aW9uX2ZhY3RvciIsDQogICAgICAgICAgICAgICAgICAgICAgICAiUmVsYXRpb25zaGlwIER1cmF0aW9uIiwNCiAgICAgICAgICAgICAgICAgICAgICAgIGlmZWxzZShjb25kaXRpb24gJWNvbnRhaW5zJSAiYl9uZXRfaW5jb21lIiwNCiAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAiSW5jb21lIiwNCiAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICBOQSkpLA0KICAgICAgICAgY29uZGl0aW9uID0gaWZlbHNlKGNvbmRpdGlvbiA9PSAiYl9hZ2UiLCAiQWdlIiwNCiAgICAgICAgICAgICAgICBpZmVsc2UoY29uZGl0aW9uID09ICJiX25ldF9pbmNvbWVldXJvXzUwMF8xMDAwIiwgIjUwMCBcVTIwMTMgMSwwMDAgRXVybyBtb250aGx5IiwNCiAgICAgICAgICAgICAgICBpZmVsc2UoY29uZGl0aW9uID09ICJiX25ldF9pbmNvbWVldXJvXzEwMDBfMjAwMCIsICIxLDAwMCBcVTIwMTMgMiwwMDAgRXVybyBtb250aGx5IiwNCiAgICAgICAgICAgICAgICBpZmVsc2UoY29uZGl0aW9uID09ICJiX25ldF9pbmNvbWVldXJvXzIwMDBfMzAwMCIsICIyLDAwMCBcVTIwMTMgMywwMDAgRXVybyBtb250aGx5IiwNCiAgICAgICAgICAgICAgICBpZmVsc2UoY29uZGl0aW9uID09ICJiX25ldF9pbmNvbWVldXJvX2d0XzMwMDAiLCAiPjMsMDAwIEV1cm8gbW9udGhseSIsDQogICAgICAgICAgICAgICAgaWZlbHNlKGNvbmRpdGlvbiA9PSAiYl9uZXRfaW5jb21lZG9udF90ZWxsIiwgImRvIG5vdCBkaXNjbG9zZSIsDQogICAgICAgICAgICAgICAgaWZlbHNlKGNvbmRpdGlvbiA9PSAiYl9yZWxhdGlvbnNoaXBfZHVyYXRpb25fZmFjdG9yUGFydG5lcmVkX3VwdG8xMm1vbnRocyIsDQogICAgICAgICAgICAgICAgICAgICAgICIwIFxVMjAxMyAxMiBtb250aHMiLA0KICAgICAgICAgICAgICAgIGlmZWxzZShjb25kaXRpb24gPT0gImJfcmVsYXRpb25zaGlwX2R1cmF0aW9uX2ZhY3RvclBhcnRuZXJlZF91cHRvMjhtb250aHMiLA0KICAgICAgICAgICAgICAgICAgICAgICAiMTMgXFUyMDEzIDI4IG1vbnRocyIsDQogICAgICAgICAgICAgICAgaWZlbHNlKGNvbmRpdGlvbiA9PSAiYl9yZWxhdGlvbnNoaXBfZHVyYXRpb25fZmFjdG9yUGFydG5lcmVkX3VwdG81Mm1vbnRocyIsDQogICAgICAgICAgICAgICAgICAgICAgICIyOSBcVTIwMTMgNTIgbW9udGhzIiwNCiAgICAgICAgICAgICAgICBpZmVsc2UoY29uZGl0aW9uID09ICJiX3JlbGF0aW9uc2hpcF9kdXJhdGlvbl9mYWN0b3JQYXJ0bmVyZWRfbW9yZXRoYW41Mm1vbnRocyIsDQogICAgICAgICAgICAgICAgICAgICAgICI+NTIgbW9udGhzIiwNCiAgICAgICAgICAgICAgICBpZmVsc2UoY29uZGl0aW9uID09ICJiX2VkdWNhdGlvbl95ZWFycyIsICJZZWFycyBvZiBFZHVjYXRpb24iLA0KICAgICAgICAgICAgICAgIGlmZWxzZShjb25kaXRpb24gPT0gImJfYmZpX2V4dHJhIiwgIkV4dHJhdmVyc2lvbiIsDQogICAgICAgICAgICAgICAgaWZlbHNlKGNvbmRpdGlvbiA9PSAiYl9iZmlfbmV1cm8iLCAiTmV1cm90aWNpc20iLA0KICAgICAgICAgICAgICAgIGlmZWxzZShjb25kaXRpb24gPT0gImJfYmZpX2FncmVlIiwgIkFncmVlYWJsZW5lc3MiLA0KICAgICAgICAgICAgICAgIGlmZWxzZShjb25kaXRpb24gPT0gImJfYmZpX2NvbnNjIiwgIkNvbnNjaWVudGlvdXNuZXNzIiwNCiAgICAgICAgICAgICAgICBpZmVsc2UoY29uZGl0aW9uID09ICJiX2JmaV9vcGVuIiwgIk9wZW5uZXNzIiwNCiAgICAgICAgICAgICAgICBpZmVsc2UoY29uZGl0aW9uID09ICJiX3JlbGlnaW9zaXR5IiwgIlJlbGlnaW9zaXR5IiwNCiAgICAgICAgICAgICAgICAgICAgICAgY29uZGl0aW9uKSkpKSkpKSkpKSkpKSkpKSksDQogICAgICAgICBncm91cCA9IGlmZWxzZShpcy5uYShncm91cCksIGNvbmRpdGlvbiwgZ3JvdXApLA0KICAgICAgICAgY29uZGl0aW9uID0gZmFjdG9yKGNvbmRpdGlvbiwgbGV2ZWxzID0gcmV2KGMoIkFnZSIsDQogICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgIjUwMCBcVTIwMTMgMSwwMDAgRXVybyBtb250aGx5IiwgIjEsMDAwIFxVMjAxMyAyLDAwMCBFdXJvIG1vbnRobHkiLA0KICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICIyLDAwMCBcVTIwMTMgMywwMDAgRXVybyBtb250aGx5IiwgIj4zLDAwMCBFdXJvIG1vbnRobHkiLCAiZG8gbm90IGRpc2Nsb3NlIiwNCiAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAiMTMgXFUyMDEzIDI4IG1vbnRocyIsICIyOSBcVTIwMTMgNTIgbW9udGhzIiwNCiAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAiPjUyIG1vbnRocyIsDQogICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgIlllYXJzIG9mIEVkdWNhdGlvbiIsDQogICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgIkV4dHJhdmVyc2lvbiIsICJOZXVyb3RpY2lzbSIsICJBZ3JlZWFibGVuZXNzIiwNCiAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgIkNvbnNjaWVudGlvdXNuZXNzIiwiT3Blbm5lc3MiLCJSZWxpZ2lvc2l0eSIpKSksDQogICAgICAgICBncm91cCA9IGZhY3Rvcihncm91cCwgbGV2ZWxzID0gYygiQWdlIiwgIkluY29tZSIsICJSZWxhdGlvbnNoaXAgRHVyYXRpb24iLA0KICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgDQogICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAiWWVhcnMgb2YgRWR1Y2F0aW9uIiwNCiAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICJFeHRyYXZlcnNpb24iLCAiTmV1cm90aWNpc20iLCAiQWdyZWVhYmxlbmVzcyIsDQogICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICJDb25zY2llbnRpb3VzbmVzcyIsIk9wZW5uZXNzIiwiUmVsaWdpb3NpdHkiKSkpICU+JQ0KICBnZ3Bsb3QoYWVzKHkgPSBjb25kaXRpb24sIHggPSBjb25kaXRpb25fbWVhbiwgY29sb3IgPSBncm91cCkpICsNCiAgc3RhdF9oYWxmZXllKC53aWR0aCA9IC45MCkgKw0KICBnZW9tX3ZsaW5lKHhpbnRlcmNlcHQgPSAwLCBsaW5ldHlwZSA9ICJkb3R0ZWQiLCBzaXplID0gMSkgKw0KICBhcGF0aGVtZSArDQogIHRoZW1lKGxlZ2VuZC50aXRsZSA9IGVsZW1lbnRfYmxhbmsoKSkgKw0KICBsYWJzKHggPSAiRWZmZWN0IFNpemUgRXN0aW1hdGVzIiwgeSA9ICJQcmVkaWN0b3JzIikgKw0KICBzY2FsZV94X2NvbnRpbnVvdXMobGltaXRzID0gYygtMywgMyksDQogICAgICAgICAgICAgICAgICAgICBicmVha3MgPSBjKC0zOjMpKSArDQogIHNjYWxlX2NvbG9yX21hbnVhbCh2YWx1ZXMgPSBjKCIjMUI5RTc3IiwgIiNEOTVGMDIiLCAiIzc1NzBCMyIsICIjRTcyOThBIiwgIiM2NkE2MUUiLA0KICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAiI0U2QUIwMiIsICIjQTY3NjFEIiwgIiMwMDcyQjIiLCAiI0NDNzlBNyIsICIjNTZCNEU5IikpICsNCiAgZ3VpZGVzKGNvbG9yID0gZ3VpZGVfbGVnZW5kKG92ZXJyaWRlLmFlcyA9IGxpc3Qoc2l6ZSA9IDUpKSkrDQogIGdndGl0bGUoZXhwcmVzc2lvbihwYXN0ZShib2xkKCJDb21wbGV4IE1vZGVsIEluaXRpYWxseSBIQyBVc2VyICgiKSwNCiAgICAgICAgICAgICAgICAgICAgICAgICAgIGJvbGRpdGFsaWMoIm4iKSwNCiAgICAgICAgICAgICAgICAgICAgICAgICAgIGJvbGQoIiA9IDM5MCkiKSkpKSArDQogIHRoZW1lKHBsb3QudGl0bGUgPSBlbGVtZW50X3RleHQoaGp1c3QgPSAtMC4zNSkpDQptX3NlbGVjdGlvbl9jb25ncnVlbnRfY29tcGxleF9vbmNlSENfcGxvdA0KDQptX3NlbGVjdGlvbl9jb25ncnVlbnRfY29tcGxleF9vbmNlbm9uSENfcGxvdCA9IG1fc2VsZWN0aW9uX2NvbmdydWVudF9jb21wbGV4X29uY2Vub25IQyAlPiUNCiAgc3ByZWFkX2RyYXdzKGJfYWdlLA0KICAgICAgICAgICAgICAgYl9uZXRfaW5jb21lZXVyb181MDBfMTAwMCwgYl9uZXRfaW5jb21lZXVyb18xMDAwXzIwMDAsIA0KICAgICAgICAgICAgICAgYl9uZXRfaW5jb21lZXVyb18yMDAwXzMwMDAsIGJfbmV0X2luY29tZWV1cm9fZ3RfMzAwMCwgYl9uZXRfaW5jb21lZG9udF90ZWxsLA0KICAgICAgICAgICAgICAgYl9yZWxhdGlvbnNoaXBfZHVyYXRpb25fZmFjdG9yUGFydG5lcmVkX3VwdG8yOG1vbnRocywNCiAgICAgICAgICAgICAgIGJfcmVsYXRpb25zaGlwX2R1cmF0aW9uX2ZhY3RvclBhcnRuZXJlZF91cHRvNTJtb250aHMsDQogICAgICAgICAgICAgICBiX3JlbGF0aW9uc2hpcF9kdXJhdGlvbl9mYWN0b3JQYXJ0bmVyZWRfbW9yZXRoYW41Mm1vbnRocywNCiAgICAgICAgICAgICAgIGJfZWR1Y2F0aW9uX3llYXJzLA0KICAgICAgICAgICAgICAgYl9iZmlfZXh0cmEsIGJfYmZpX25ldXJvLCBiX2JmaV9hZ3JlZSwgYl9iZmlfY29uc2MsIGJfYmZpX29wZW4sIA0KICAgICAgICAgICAgICAgYl9yZWxpZ2lvc2l0eSkgJT4lDQogIHBpdm90X2xvbmdlcihjb2xzID0gYyhiX2FnZSwNCiAgICAgICAgICAgICAgIGJfbmV0X2luY29tZWV1cm9fNTAwXzEwMDAsIGJfbmV0X2luY29tZWV1cm9fMTAwMF8yMDAwLCANCiAgICAgICAgICAgICAgIGJfbmV0X2luY29tZWV1cm9fMjAwMF8zMDAwLCBiX25ldF9pbmNvbWVldXJvX2d0XzMwMDAsIGJfbmV0X2luY29tZWRvbnRfdGVsbCwNCiAgICAgICAgICAgICAgIGJfcmVsYXRpb25zaGlwX2R1cmF0aW9uX2ZhY3RvclBhcnRuZXJlZF91cHRvMjhtb250aHMsDQogICAgICAgICAgICAgICBiX3JlbGF0aW9uc2hpcF9kdXJhdGlvbl9mYWN0b3JQYXJ0bmVyZWRfdXB0bzUybW9udGhzLA0KICAgICAgICAgICAgICAgYl9yZWxhdGlvbnNoaXBfZHVyYXRpb25fZmFjdG9yUGFydG5lcmVkX21vcmV0aGFuNTJtb250aHMsDQogICAgICAgICAgICAgICBiX2VkdWNhdGlvbl95ZWFycywNCiAgICAgICAgICAgICAgIGJfYmZpX2V4dHJhLCBiX2JmaV9uZXVybywgYl9iZmlfYWdyZWUsIGJfYmZpX2NvbnNjLCBiX2JmaV9vcGVuLCANCiAgICAgICAgICAgICAgIGJfcmVsaWdpb3NpdHkpLA0KICAgICAgICAgICAgICAgbmFtZXNfdG8gPSAiY29uZGl0aW9uIiwNCiAgICAgICAgICAgICAgIHZhbHVlc190byA9ICJyX2NvbmRpdGlvbiIpICU+JQ0KICBtdXRhdGUoY29uZGl0aW9uX21lYW4gPSByX2NvbmRpdGlvbiwNCiAgICAgICAgIGdyb3VwID0gaWZlbHNlKGNvbmRpdGlvbiAlY29udGFpbnMlICJiX3JlbGF0aW9uc2hpcF9kdXJhdGlvbl9mYWN0b3IiLA0KICAgICAgICAgICAgICAgICAgICAgICAgIlJlbGF0aW9uc2hpcCBEdXJhdGlvbiIsDQogICAgICAgICAgICAgICAgICAgICAgICBpZmVsc2UoY29uZGl0aW9uICVjb250YWlucyUgImJfbmV0X2luY29tZSIsDQogICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgIkluY29tZSIsDQogICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgTkEpKSwNCiAgICAgICAgIGNvbmRpdGlvbiA9IGlmZWxzZShjb25kaXRpb24gPT0gImJfYWdlIiwgIkFnZSIsDQogICAgICAgICAgICAgICAgaWZlbHNlKGNvbmRpdGlvbiA9PSAiYl9uZXRfaW5jb21lZXVyb181MDBfMTAwMCIsICI1MDAgXFUyMDEzIDEsMDAwIEV1cm8gbW9udGhseSIsDQogICAgICAgICAgICAgICAgaWZlbHNlKGNvbmRpdGlvbiA9PSAiYl9uZXRfaW5jb21lZXVyb18xMDAwXzIwMDAiLCAiMSwwMDAgXFUyMDEzIDIsMDAwIEV1cm8gbW9udGhseSIsDQogICAgICAgICAgICAgICAgaWZlbHNlKGNvbmRpdGlvbiA9PSAiYl9uZXRfaW5jb21lZXVyb18yMDAwXzMwMDAiLCAiMiwwMDAgXFUyMDEzIDMsMDAwIEV1cm8gbW9udGhseSIsDQogICAgICAgICAgICAgICAgaWZlbHNlKGNvbmRpdGlvbiA9PSAiYl9uZXRfaW5jb21lZXVyb19ndF8zMDAwIiwgIj4zLDAwMCBFdXJvIG1vbnRobHkiLA0KICAgICAgICAgICAgICAgIGlmZWxzZShjb25kaXRpb24gPT0gImJfbmV0X2luY29tZWRvbnRfdGVsbCIsICJkbyBub3QgZGlzY2xvc2UiLA0KICAgICAgICAgICAgICAgIGlmZWxzZShjb25kaXRpb24gPT0gImJfcmVsYXRpb25zaGlwX2R1cmF0aW9uX2ZhY3RvclBhcnRuZXJlZF91cHRvMTJtb250aHMiLA0KICAgICAgICAgICAgICAgICAgICAgICAiMCBcVTIwMTMgMTIgbW9udGhzIiwNCiAgICAgICAgICAgICAgICBpZmVsc2UoY29uZGl0aW9uID09ICJiX3JlbGF0aW9uc2hpcF9kdXJhdGlvbl9mYWN0b3JQYXJ0bmVyZWRfdXB0bzI4bW9udGhzIiwNCiAgICAgICAgICAgICAgICAgICAgICAgIjEzIFxVMjAxMyAyOCBtb250aHMiLA0KICAgICAgICAgICAgICAgIGlmZWxzZShjb25kaXRpb24gPT0gImJfcmVsYXRpb25zaGlwX2R1cmF0aW9uX2ZhY3RvclBhcnRuZXJlZF91cHRvNTJtb250aHMiLA0KICAgICAgICAgICAgICAgICAgICAgICAiMjkgXFUyMDEzIDUyIG1vbnRocyIsDQogICAgICAgICAgICAgICAgaWZlbHNlKGNvbmRpdGlvbiA9PSAiYl9yZWxhdGlvbnNoaXBfZHVyYXRpb25fZmFjdG9yUGFydG5lcmVkX21vcmV0aGFuNTJtb250aHMiLA0KICAgICAgICAgICAgICAgICAgICAgICAiPjUyIG1vbnRocyIsDQogICAgICAgICAgICAgICAgaWZlbHNlKGNvbmRpdGlvbiA9PSAiYl9lZHVjYXRpb25feWVhcnMiLCAiWWVhcnMgb2YgRWR1Y2F0aW9uIiwNCiAgICAgICAgICAgICAgICBpZmVsc2UoY29uZGl0aW9uID09ICJiX2JmaV9leHRyYSIsICJFeHRyYXZlcnNpb24iLA0KICAgICAgICAgICAgICAgIGlmZWxzZShjb25kaXRpb24gPT0gImJfYmZpX25ldXJvIiwgIk5ldXJvdGljaXNtIiwNCiAgICAgICAgICAgICAgICBpZmVsc2UoY29uZGl0aW9uID09ICJiX2JmaV9hZ3JlZSIsICJBZ3JlZWFibGVuZXNzIiwNCiAgICAgICAgICAgICAgICBpZmVsc2UoY29uZGl0aW9uID09ICJiX2JmaV9jb25zYyIsICJDb25zY2llbnRpb3VzbmVzcyIsDQogICAgICAgICAgICAgICAgaWZlbHNlKGNvbmRpdGlvbiA9PSAiYl9iZmlfb3BlbiIsICJPcGVubmVzcyIsDQogICAgICAgICAgICAgICAgaWZlbHNlKGNvbmRpdGlvbiA9PSAiYl9yZWxpZ2lvc2l0eSIsICJSZWxpZ2lvc2l0eSIsDQogICAgICAgICAgICAgICAgICAgICAgIGNvbmRpdGlvbikpKSkpKSkpKSkpKSkpKSkpLA0KICAgICAgICAgZ3JvdXAgPSBpZmVsc2UoaXMubmEoZ3JvdXApLCBjb25kaXRpb24sIGdyb3VwKSwNCiAgICAgICAgIGNvbmRpdGlvbiA9IGZhY3Rvcihjb25kaXRpb24sIGxldmVscyA9IHJldihjKCJBZ2UiLA0KICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICI1MDAgXFUyMDEzIDEsMDAwIEV1cm8gbW9udGhseSIsICIxLDAwMCBcVTIwMTMgMiwwMDAgRXVybyBtb250aGx5IiwNCiAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAiMiwwMDAgXFUyMDEzIDMsMDAwIEV1cm8gbW9udGhseSIsICI+MywwMDAgRXVybyBtb250aGx5IiwgImRvIG5vdCBkaXNjbG9zZSIsDQogICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgIjEzIFxVMjAxMyAyOCBtb250aHMiLCAiMjkgXFUyMDEzIDUyIG1vbnRocyIsDQogICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgIj41MiBtb250aHMiLA0KICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICJZZWFycyBvZiBFZHVjYXRpb24iLA0KICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICJFeHRyYXZlcnNpb24iLCAiTmV1cm90aWNpc20iLCAiQWdyZWVhYmxlbmVzcyIsDQogICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICJDb25zY2llbnRpb3VzbmVzcyIsIk9wZW5uZXNzIiwiUmVsaWdpb3NpdHkiKSkpLA0KICAgICAgICAgZ3JvdXAgPSBmYWN0b3IoZ3JvdXAsIGxldmVscyA9IGMoIkFnZSIsICJJbmNvbWUiLCAiUmVsYXRpb25zaGlwIER1cmF0aW9uIiwNCiAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgIA0KICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgIlllYXJzIG9mIEVkdWNhdGlvbiIsDQogICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAiRXh0cmF2ZXJzaW9uIiwgIk5ldXJvdGljaXNtIiwgIkFncmVlYWJsZW5lc3MiLA0KICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAiQ29uc2NpZW50aW91c25lc3MiLCJPcGVubmVzcyIsIlJlbGlnaW9zaXR5IikpKSAlPiUNCiAgZ2dwbG90KGFlcyh5ID0gY29uZGl0aW9uLCB4ID0gY29uZGl0aW9uX21lYW4sIGNvbG9yID0gZ3JvdXApKSArDQogIHN0YXRfaGFsZmV5ZSgud2lkdGggPSAuOTApICsNCiAgZ2VvbV92bGluZSh4aW50ZXJjZXB0ID0gMCwgbGluZXR5cGUgPSAiZG90dGVkIiwgc2l6ZSA9IDEpICsNCiAgYXBhdGhlbWUgKw0KICB0aGVtZShsZWdlbmQudGl0bGUgPSBlbGVtZW50X2JsYW5rKCkpICsNCiAgbGFicyh4ID0gIkVmZmVjdCBTaXplIEVzdGltYXRlcyIsIHkgPSAiUHJlZGljdG9ycyIpICsNCiAgc2NhbGVfeF9jb250aW51b3VzKGxpbWl0cyA9IGMoLTMsIDMpLA0KICAgICAgICAgICAgICAgICAgICAgYnJlYWtzID0gYygtMzozKSkgKw0KICBzY2FsZV9jb2xvcl9tYW51YWwodmFsdWVzID0gYygiIzFCOUU3NyIsICIjRDk1RjAyIiwgIiM3NTcwQjMiLCAiI0U3Mjk4QSIsICIjNjZBNjFFIiwNCiAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgIiNFNkFCMDIiLCAiI0E2NzYxRCIsICIjMDA3MkIyIiwgIiNDQzc5QTciLCAiIzU2QjRFOSIpKSArDQogIGd1aWRlcyhjb2xvciA9IGd1aWRlX2xlZ2VuZChvdmVycmlkZS5hZXMgPSBsaXN0KHNpemUgPSA1KSkpKw0KICBnZ3RpdGxlKGV4cHJlc3Npb24ocGFzdGUoYm9sZCgiQ29tcGxleCBNb2RlbCBJbml0aWFsbHkgTm9uLUhDIFVzZXIgKCIpLA0KICAgICAgICAgICAgICAgICAgICAgICAgICAgYm9sZGl0YWxpYygibiIpLA0KICAgICAgICAgICAgICAgICAgICAgICAgICAgYm9sZCgiID0gMzg0KSIpKSkpICsNCiAgdGhlbWUocGxvdC50aXRsZSA9IGVsZW1lbnRfdGV4dChoanVzdCA9IC0wLjM1KSkNCg0KbV9zZWxlY3Rpb25fY29uZ3J1ZW50X2NvbXBsZXhfb25jZW5vbkhDX3Bsb3QNCmBgYA0KDQojIyMjIENvbWJpbmUgUGxvdHMNCmBgYHtyfQ0Kc2VsZWN0aW9uX2NvbmdydWVudCA9DQogIGdnYXJyYW5nZShtX3NlbGVjdGlvbl9jb25ncnVlbnRfc2ltcGxlX29uY2VIQ19wbG90ICsNCiAgICAgICAgICAgICAgdGhlbWUobGVnZW5kLnBvc2l0aW9uID0gIm5vbmUiLCBwbG90Lm1hcmdpbiA9IG1hcmdpbigxLDAsMCwwLCJjbSIpKSwNCiAgICAgICAgICAgIG1fc2VsZWN0aW9uX2NvbmdydWVudF9zaW1wbGVfb25jZW5vbkhDX3Bsb3QgKw0KICAgICAgICAgICAgICB0aGVtZShsZWdlbmQucG9zaXRpb24gPSBjKDAuODUsMC41KSwgcGxvdC5tYXJnaW4gPSBtYXJnaW4oMSwwLDAsMCwiY20iKSksDQogICAgICAgICAgICBtX3NlbGVjdGlvbl9jb25ncnVlbnRfY29tcGxleF9vbmNlSENfcGxvdCArDQogICAgICAgICAgICAgIHRoZW1lKGxlZ2VuZC5wb3NpdGlvbiA9ICJub25lIiwgcGxvdC5tYXJnaW4gPSBtYXJnaW4oMSwwLDAsMCwiY20iKSksDQogICAgICAgICAgICBtX3NlbGVjdGlvbl9jb25ncnVlbnRfY29tcGxleF9vbmNlbm9uSENfcGxvdCAgKw0KICAgICAgICAgICAgICB0aGVtZShsZWdlbmQucG9zaXRpb24gPSBjKDAuODUsMC41KSwgcGxvdC5tYXJnaW4gPSBtYXJnaW4oMSwwLDAsMCwiY20iKSksDQogICAgICAgICAgaGp1c3QgPSAtMC4xLCB2anVzdCA9IDEuMSwNCiAgICAgICAgICBuY29sID0gMiwgbnJvdyA9IDIsDQogICAgICAgICAgYWxpZ24gPSAidiIpDQoNCnNlbGVjdGlvbl9jb25ncnVlbnQNCmpwZWcoJ0VmZmVjdCBzaXplIGVzdGltYXRlcyBmb3IgbW9kZWxzIHdpdGggY29uZ3J1ZW5jeSBpbiBjb250cmFjZXB0aXZlIG1ldGhvZCBhcyBhbiBvdXRjb21lLmpwZycsIA0KICAgICB3aWR0aCA9IDE2MDAsIGhlaWdodCA9IDcyMCwgcXVhbGl0eSA9IDEwMCkNCnNlbGVjdGlvbl9jb25ncnVlbnQNCmRldi5vZmYoKQ0KYGBgDQoNCg0KIyMgRWZmZWN0cyBvZiBIb3Jtb25hbCBDb250cmFjZXB0aXZlcyB7LnRhYnNldH0NCiMjIyBVbmNvbnRyb2xsZWQNCiMjIyMgTW9kZWxzDQpgYGB7cn0NCm1faGNfYXRyciA9IGJybShhdHRyYWN0aXZlbmVzc19wYXJ0bmVyIH4gY29udHJhY2VwdGlvbl9ob3Jtb25hbCwNCiAgICAgICAgICAgICAgICBkYXRhID0gZGF0YSwgZmFtaWx5ID0gZ2F1c3NpYW4oKSwNCiAgICAgICAgICAgICAgICBmaWxlID0gIm1faGNfYXRyciIpDQptX2hjX3JlbHNhdCA9IGJybShyZWxhdGlvbnNoaXBfc2F0aXNmYWN0aW9uIH4gY29udHJhY2VwdGlvbl9ob3Jtb25hbCwNCiAgICAgICAgICAgICAgICBkYXRhID0gZGF0YSwgZmFtaWx5ID0gZ2F1c3NpYW4oKSwNCiAgICAgICAgICAgICAgICBmaWxlID0gIm1faGNfcmVsc2F0IikNCm1faGNfc2V4c2F0ID0gYnJtKHNhdGlzZmFjdGlvbl9zZXh1YWxfaW50ZXJjb3Vyc2UgfiBjb250cmFjZXB0aW9uX2hvcm1vbmFsLA0KICAgICAgICAgICAgICAgIGRhdGEgPSBkYXRhLCBmYW1pbHkgPSBnYXVzc2lhbigpLA0KICAgICAgICAgICAgICAgIGZpbGUgPSAibV9oY19zZXhzYXQiKQ0KbV9oY19saWJpZG8gPSBicm0oZGlhcnlfbGliaWRvX21lYW4gfiBjb250cmFjZXB0aW9uX2hvcm1vbmFsLA0KICAgICAgICAgICAgICAgIGRhdGEgPSBkYXRhLCBmYW1pbHkgPSBnYXVzc2lhbigpLA0KICAgICAgICAgICAgICAgIGZpbGUgPSAibV9oY19saWJpZG8iKQ0KbV9oY19zZXhmcmVxcGVuID0gYnJtKGRpYXJ5X3NleF9hY3RpdmVfc2V4X3N1bSB+DQogICAgICAgICAgICAgICAgICAgICAgICBvZmZzZXQobG9nKG51bWJlcl9vZl9kYXlzKSkgKw0KICAgICAgICAgICAgICAgICAgICAgICAgY29udHJhY2VwdGlvbl9ob3Jtb25hbCwNCiAgICAgICAgICAgICAgICBkYXRhID0gZGF0YSwgZmFtaWx5ID0gcG9pc3NvbigpLA0KICAgICAgICAgICAgICAgIGZpbGUgPSAibXJvYnVzdF9oY19zZXhmcmVxcGVuIikNCm1faGNfbWFzZnJlcSA9IGJybShkaWFyeV9tYXN0dXJiYXRpb25fc3VtIH4gb2Zmc2V0KGxvZyhudW1iZXJfb2ZfZGF5cykpICsNCiAgICAgICAgICAgICAgICAgICAgIGNvbnRyYWNlcHRpb25faG9ybW9uYWwsDQogICAgICAgICAgICAgICAgZGF0YSA9IGRhdGEsIGZhbWlseSA9IHBvaXNzb24oKSwNCiAgICAgICAgICAgICAgICBmaWxlID0gIm1yb2J1c3RfaGNfbWFzZnJlcSIpDQpgYGANCg0KIyMjIyBGb3Jlc3QgUGxvdCBmb3IgRWZmZWN0IFNpemVzIHsuYWN0aXZlfQ0KYGBge3J9DQptb2RlbHMgPSByYmluZCgNCiAgbV9oY19hdHJyICU+JQ0KICAgIHNwcmVhZF9kcmF3cyhiX2NvbnRyYWNlcHRpb25faG9ybW9uYWx5ZXMpICU+JQ0KICAgIHBpdm90X2xvbmdlcihjb2xzID0gYyhiX2NvbnRyYWNlcHRpb25faG9ybW9uYWx5ZXMpLA0KICAgICAgICAgICAgICAgICBuYW1lc190byA9ICJjb25kaXRpb24iLA0KICAgICAgICAgICAgICAgICB2YWx1ZXNfdG8gPSAicl9jb25kaXRpb24iKSAlPiUNCiAgICBtdXRhdGUobW9kZWwgPSAiUGVyY2VpdmVkIFBhcnRuZXIgQXR0cmFjdGl2ZW5lc3NcblJPUEUgPSBbLTAuMDcsIDAuMDddIiwNCiAgICAgICAgICAgUk9QRSA9IDAuMDcpLA0KICBtX2hjX3JlbHNhdCAlPiUNCiAgICBzcHJlYWRfZHJhd3MoYl9jb250cmFjZXB0aW9uX2hvcm1vbmFseWVzKSAlPiUNCiAgICBwaXZvdF9sb25nZXIoY29scyA9IGMoYl9jb250cmFjZXB0aW9uX2hvcm1vbmFseWVzKSwNCiAgICAgICAgICAgICAgICAgbmFtZXNfdG8gPSAiY29uZGl0aW9uIiwNCiAgICAgICAgICAgICAgICAgdmFsdWVzX3RvID0gInJfY29uZGl0aW9uIikgJT4lDQogICAgbXV0YXRlKG1vZGVsID0gIlJlbGF0aW9uc2hpcCBTYXRpc2ZhY3Rpb25cblJPUEUgPSBbLTAuMDQsIDAuMDRdIiwNCiAgICAgICAgICAgUk9QRSA9IDAuMDQpLA0KICBtX2hjX3NleHNhdCAlPiUNCiAgICBzcHJlYWRfZHJhd3MoYl9jb250cmFjZXB0aW9uX2hvcm1vbmFseWVzKSAlPiUNCiAgICBwaXZvdF9sb25nZXIoY29scyA9IGMoYl9jb250cmFjZXB0aW9uX2hvcm1vbmFseWVzKSwNCiAgICAgICAgICAgICAgICAgbmFtZXNfdG8gPSAiY29uZGl0aW9uIiwNCiAgICAgICAgICAgICAgICAgdmFsdWVzX3RvID0gInJfY29uZGl0aW9uIikgJT4lDQogICAgbXV0YXRlKG1vZGVsID0gIlNleHVhbCBTYXRpc2ZhY3Rpb25cblJPUEUgPSBbLTAuMTEsIDAuMTFdIiwNCiAgICAgICAgICAgUk9QRSA9IDAuMTEpLA0KICBtX2hjX2xpYmlkbyAlPiUNCiAgICBzcHJlYWRfZHJhd3MoYl9jb250cmFjZXB0aW9uX2hvcm1vbmFseWVzKSAlPiUNCiAgICBwaXZvdF9sb25nZXIoY29scyA9IGMoYl9jb250cmFjZXB0aW9uX2hvcm1vbmFseWVzKSwNCiAgICAgICAgICAgICAgICAgbmFtZXNfdG8gPSAiY29uZGl0aW9uIiwNCiAgICAgICAgICAgICAgICAgdmFsdWVzX3RvID0gInJfY29uZGl0aW9uIikgJT4lDQogICAgbXV0YXRlKG1vZGVsID0gIkxpYmlkb1xuUk9QRSA9IFstMC4wNiwgMC4wNl0iLA0KICAgICAgICAgICBST1BFID0gMC4wNiksDQogIG1faGNfc2V4ZnJlcXBlbiAlPiUNCiAgICBzcHJlYWRfZHJhd3MoYl9jb250cmFjZXB0aW9uX2hvcm1vbmFseWVzKSAlPiUNCiAgICBwaXZvdF9sb25nZXIoY29scyA9IGMoYl9jb250cmFjZXB0aW9uX2hvcm1vbmFseWVzKSwNCiAgICAgICAgICAgICAgICAgbmFtZXNfdG8gPSAiY29uZGl0aW9uIiwNCiAgICAgICAgICAgICAgICAgdmFsdWVzX3RvID0gInJfY29uZGl0aW9uIikgJT4lDQogICAgbXV0YXRlKG1vZGVsID0gIkZyZXF1ZW5jeSBvZiBWYWdpbmFsIEludGVyY291cnNlXG5ST1BFID0gWy0wLjA1LCAwLjA1XSIsDQogICAgICAgICAgIFJPUEUgPSAwLjA1KSwNCiAgbV9oY19tYXNmcmVxICU+JQ0KICAgIHNwcmVhZF9kcmF3cyhiX2NvbnRyYWNlcHRpb25faG9ybW9uYWx5ZXMpICU+JQ0KICAgIHBpdm90X2xvbmdlcihjb2xzID0gYyhiX2NvbnRyYWNlcHRpb25faG9ybW9uYWx5ZXMpLA0KICAgICAgICAgICAgICAgICBuYW1lc190byA9ICJjb25kaXRpb24iLA0KICAgICAgICAgICAgICAgICB2YWx1ZXNfdG8gPSAicl9jb25kaXRpb24iKSAlPiUNCiAgICBtdXRhdGUobW9kZWwgPSAiRnJlcXVlbmN5IG9mIE1hc3R1cmJhdGlvblxuUk9QRSA9IFstMC4wNSwgMC4wNV0iLA0KICAgICAgICAgICBST1BFID0gMC4wNSkpDQoNCmRhdGExID0gbW9kZWxzICU+JQ0KICBtdXRhdGUobW9kZWwgPSBmYWN0b3IobW9kZWwsDQogICAgICAgICAgICAgICAgICAgbGV2ZWxzID0gYygiRnJlcXVlbmN5IG9mIE1hc3R1cmJhdGlvblxuUk9QRSA9IFstMC4wNSwgMC4wNV0iLA0KICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgIkZyZXF1ZW5jeSBvZiBWYWdpbmFsIEludGVyY291cnNlXG5ST1BFID0gWy0wLjA1LCAwLjA1XSIsDQogICAgICAgICAgICAgICAgICAgICAgICAgICAgICAiTGliaWRvXG5ST1BFID0gWy0wLjA2LCAwLjA2XSIsDQogICAgICAgICAgICAgICAgICAgICAgICAgICAgICAiU2V4dWFsIFNhdGlzZmFjdGlvblxuUk9QRSA9IFstMC4xMSwgMC4xMV0iLA0KICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgIlJlbGF0aW9uc2hpcCBTYXRpc2ZhY3Rpb25cblJPUEUgPSBbLTAuMDQsIDAuMDRdIiwNCiAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICJQZXJjZWl2ZWQgUGFydG5lciBBdHRyYWN0aXZlbmVzc1xuUk9QRSA9IFstMC4wNywgMC4wN10iKSkpDQpkYXRhMiA9IG1vZGVscyAlPiUNCiAgZ3JvdXBfYnkobW9kZWwsIFJPUEUpICU+JQ0KICBtdXRhdGUobW9kZWwgPSBmYWN0b3IobW9kZWwsDQogICAgICAgICAgICAgICAgICAgICAgICBsZXZlbHMgPSBjKCJGcmVxdWVuY3kgb2YgTWFzdHVyYmF0aW9uXG5ST1BFID0gWy0wLjA1LCAwLjA1XSIsDQogICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICJGcmVxdWVuY3kgb2YgVmFnaW5hbCBJbnRlcmNvdXJzZVxuUk9QRSA9IFstMC4wNSwgMC4wNV0iLA0KICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAiTGliaWRvXG5ST1BFID0gWy0wLjA2LCAwLjA2XSIsDQogICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICJTZXh1YWwgU2F0aXNmYWN0aW9uXG5ST1BFID0gWy0wLjExLCAwLjExXSIsDQogICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICJSZWxhdGlvbnNoaXAgU2F0aXNmYWN0aW9uXG5ST1BFID0gWy0wLjA0LCAwLjA0XSIsDQogICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICJQZXJjZWl2ZWQgUGFydG5lciBBdHRyYWN0aXZlbmVzc1xuUk9QRSA9IFstMC4wNywgMC4wN10iKSkpICU+JQ0KICBtZWFuX3FpKGNvbmRpdGlvbl9tZWFuID0gcl9jb25kaXRpb24sDQogICAgICAgICAgLndpZHRoID0gYyguOTApKSAlPiUNCiAgbXV0YXRlKHRleHQgPSBpZmVsc2UoLndpZHRoID09IC45MCwNCiAgICAgICAgICAgICAgICAgICAgICAgc3RyX2Mocm91bmQoY29uZGl0aW9uX21lYW4sIDIpLA0KICAgICAgICAgICAgICAgICAgICAgICAgICAgICAiIFsiLA0KICAgICAgICAgICAgICAgICAgICAgICAgICAgICByb3VuZCgubG93ZXIsIDIpLA0KICAgICAgICAgICAgICAgICAgICAgICAgICAgICAiOyAiLA0KICAgICAgICAgICAgICAgICAgICAgICAgICAgICByb3VuZCgudXBwZXIsIDIpLA0KICAgICAgICAgICAgICAgICAgICAgICAgICAgICAiXSIpLCBOQSksDQogICAgICAgICBzaWcgPSBpZmVsc2UoLmxvd2VyID4gUk9QRSB8IC51cHBlciA8IC1ST1BFLCBULCBGKSkNCg0KDQpoY191bmNvbnRyb2xsZWQgPSBnZ3Bsb3QoKSArDQogIHN0YXRfaGFsZmV5ZShkYXRhID0gZGF0YTEsDQogICAgICAgICAgICAgICBhZXMoeSA9IG1vZGVsLCB4ID0gcl9jb25kaXRpb24sDQogICAgICAgICAgICAgICAgICAgY29sb3IgPSAiZ3JleSIpLA0KICAgICAgICAgICAgICAgLndpZHRoID0gLjkwKSArDQogIGdlb21fdmxpbmUoeGludGVyY2VwdCA9IDAsIGxpbmV0eXBlID0gImRvdHRlZCIpICsNCiAgZ2VvbV9wb2ludGludGVydmFsKGRhdGEgPSBkYXRhMiwNCiAgICAgICAgICAgICAgICAgICAgIGFlcyh5ID0gbW9kZWwsDQogICAgICAgICAgICAgICAgICAgICAgICAgeCA9IGNvbmRpdGlvbl9tZWFuLA0KICAgICAgICAgICAgICAgICAgICAgICAgIHhtaW4gPSAubG93ZXIsDQogICAgICAgICAgICAgICAgICAgICAgICAgeG1heCA9IC51cHBlciwNCiAgICAgICAgICAgICAgICAgICAgICAgICBjb2xvciA9IHNpZykpICsNCiAgZ2VvbV9wb2ludChkYXRhID0gZGF0YTEsIGFlcyh5ID0gbW9kZWwsIHggPSBST1BFKSwgc2hhcGUgPSAifCIsIHNpemUgPSAxMCwgY29sb3IgPSAiYmxhY2siKSArDQogIGdlb21fcG9pbnQoZGF0YSA9IGRhdGExLCBhZXMoeSA9IG1vZGVsLCB4ID0gLVJPUEUpLCBzaGFwZSA9ICJ8Iiwgc2l6ZSA9IDEwLCBjb2xvciA9ICJibGFjayIpICsNCiAgZ2VvbV9wb2ludChkYXRhID0gZGF0YTEsIGFlcyh5ID0gbW9kZWwsIHggPSBST1BFKSwgc2hhcGUgPSA1OCwgc2l6ZSA9IDEwLCBjb2xvciA9ICJ3aGl0ZSIpICsNCiAgZ2VvbV9wb2ludChkYXRhID0gZGF0YTEsIGFlcyh5ID0gbW9kZWwsIHggPSAtUk9QRSksIHNoYXBlID0gNTgsIHNpemUgPSAxMCwgY29sb3IgPSAid2hpdGUiKSArDQogIGFwYXRoZW1lICsNCiAgc2NhbGVfY29sb3JfbWFudWFsKGRhdGEyLCB2YWx1ZXMgPSBjKCJza3libHVlIiwgImdyZXkiLCAicmVkIikpICsNCiAgc2NhbGVfeF9jb250aW51b3VzKGJyZWFrcyA9IGMoLTAuNSwgLTAuNCwgLTAuMywgLTAuMiwgLTAuMSwgMCwgMC4xLCAwLjIsIDAuMywgMC40LCAwLjUpLA0KICAgICAgICAgICAgICAgICAgICAgbGltaXRzID0gYygtMC41LCAwLjUpKSArDQogIGxhYnMoeCA9ICJFZmZlY3QgU2l6ZSBFc3RpbWF0ZXMgb2YgSG9ybW9uYWwgQ29udHJhY2VwdGlvbiIsIHkgPSAiT3V0Y29tZXMiKSArDQogIHRoZW1lKGxlZ2VuZC5wb3NpdGlvbiA9ICJub25lIikNCmhjX3VuY29udHJvbGxlZA0KYGBgDQoNCiMjIyBDb250cm9sbGVkDQojIyMjIE1vZGVscw0KYGBge3J9DQptX2hjX2F0cnJfY29udHJvbGxlZCA9IGJybShhdHRyYWN0aXZlbmVzc19wYXJ0bmVyIH4gY29udHJhY2VwdGlvbl9ob3Jtb25hbCArDQogICAgICAgICAgICAgICAgICAgICAgICAgICAgIGFnZSArIG5ldF9pbmNvbWUgKyByZWxhdGlvbnNoaXBfZHVyYXRpb25fZmFjdG9yICsNCiAgICAgICAgICAgICAgICAgICAgICAgICAgICAgIGVkdWNhdGlvbl95ZWFycyArDQogICAgICAgICAgICAgICAgICAgICAgICAgICAgICBiZmlfZXh0cmEgKyBiZmlfbmV1cm8gKyBiZmlfYWdyZWUgKyBiZmlfY29uc2MgKyBiZmlfb3BlbiArDQogICAgICAgICAgICAgICAgICAgICAgICAgICAgICByZWxpZ2lvc2l0eSwNCiAgICAgICAgICAgICAgICBkYXRhID0gZGF0YSwgZmFtaWx5ID0gZ2F1c3NpYW4oKSwNCiAgICAgICAgICAgICAgICBmaWxlID0gIm1faGNfYXRycl9jb250cm9sbGVkIikNCg0KbV9oY19yZWxzYXRfY29udHJvbGxlZCA9IGJybShyZWxhdGlvbnNoaXBfc2F0aXNmYWN0aW9uIH4gY29udHJhY2VwdGlvbl9ob3Jtb25hbCArDQogICAgICAgICAgICAgICAgICAgICAgICAgICAgIGFnZSArIG5ldF9pbmNvbWUgKyByZWxhdGlvbnNoaXBfZHVyYXRpb25fZmFjdG9yICsNCiAgICAgICAgICAgICAgICAgICAgICAgICAgICAgIGVkdWNhdGlvbl95ZWFycyArDQogICAgICAgICAgICAgICAgICAgICAgICAgICAgICBiZmlfZXh0cmEgKyBiZmlfbmV1cm8gKyBiZmlfYWdyZWUgKyBiZmlfY29uc2MgKyBiZmlfb3BlbiArDQogICAgICAgICAgICAgICAgICAgICAgICAgICAgICByZWxpZ2lvc2l0eSwNCiAgICAgICAgICAgICAgICBkYXRhID0gZGF0YSwgZmFtaWx5ID0gZ2F1c3NpYW4oKSwNCiAgICAgICAgICAgICAgICBmaWxlID0gIm1faGNfcmVsc2F0X2NvbnRyb2xsZWQiKQ0KDQptX2hjX3NleHNhdF9jb250cm9sbGVkID0gYnJtKHNhdGlzZmFjdGlvbl9zZXh1YWxfaW50ZXJjb3Vyc2UgfiBjb250cmFjZXB0aW9uX2hvcm1vbmFsICsNCiAgICAgICAgICAgICAgICAgICAgICAgICAgICAgYWdlICsgbmV0X2luY29tZSArIHJlbGF0aW9uc2hpcF9kdXJhdGlvbl9mYWN0b3IgKw0KICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgZWR1Y2F0aW9uX3llYXJzICsNCiAgICAgICAgICAgICAgICAgICAgICAgICAgICAgIGJmaV9leHRyYSArIGJmaV9uZXVybyArIGJmaV9hZ3JlZSArIGJmaV9jb25zYyArIGJmaV9vcGVuICsNCiAgICAgICAgICAgICAgICAgICAgICAgICAgICAgIHJlbGlnaW9zaXR5LA0KICAgICAgICAgICAgICAgIGRhdGEgPSBkYXRhLCBmYW1pbHkgPSBnYXVzc2lhbigpLA0KICAgICAgICAgICAgICAgIGZpbGUgPSAibV9oY19zZXhzYXRfY29udHJvbGxlZCIpDQoNCm1faGNfbGliaWRvX2NvbnRyb2xsZWQgPSBicm0oZGlhcnlfbGliaWRvX21lYW4gfiBjb250cmFjZXB0aW9uX2hvcm1vbmFsICsNCiAgICAgICAgICAgICAgICAgICAgICAgICAgICAgYWdlICsgbmV0X2luY29tZSArIHJlbGF0aW9uc2hpcF9kdXJhdGlvbl9mYWN0b3IgKw0KICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgZWR1Y2F0aW9uX3llYXJzICsNCiAgICAgICAgICAgICAgICAgICAgICAgICAgICAgIGJmaV9leHRyYSArIGJmaV9uZXVybyArIGJmaV9hZ3JlZSArIGJmaV9jb25zYyArIGJmaV9vcGVuICsNCiAgICAgICAgICAgICAgICAgICAgICAgICAgICAgIHJlbGlnaW9zaXR5LA0KICAgICAgICAgICAgICAgIGRhdGEgPSBkYXRhLCBmYW1pbHkgPSBnYXVzc2lhbigpLA0KICAgICAgICAgICAgICAgIGZpbGUgPSAibV9oY19saWJpZG9fY29udHJvbGxlZCIpDQoNCm1faGNfc2V4ZnJlcXBlbl9jb250cm9sbGVkID0gYnJtKGRpYXJ5X3NleF9hY3RpdmVfc2V4X3N1bSB+DQogICAgICAgICAgICAgICAgICAgICAgICBvZmZzZXQobG9nKG51bWJlcl9vZl9kYXlzKSkgKw0KICAgICAgICAgICAgICAgICAgICAgICAgY29udHJhY2VwdGlvbl9ob3Jtb25hbCArDQogICAgICAgICAgICAgICAgICAgICAgICBhZ2UgKyBuZXRfaW5jb21lICsgcmVsYXRpb25zaGlwX2R1cmF0aW9uX2ZhY3RvciArDQogICAgICAgICAgICAgICAgICAgICAgICAgICAgICBlZHVjYXRpb25feWVhcnMgKw0KICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgYmZpX2V4dHJhICsgYmZpX25ldXJvICsgYmZpX2FncmVlICsgYmZpX2NvbnNjICsgYmZpX29wZW4gKw0KICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgcmVsaWdpb3NpdHksDQogICAgICAgICAgICAgICAgZGF0YSA9IGRhdGEsIGZhbWlseSA9IHBvaXNzb24oKSwNCiAgICAgICAgICAgICAgICBmaWxlID0gIm1yb2J1c3RfaGNfc2V4ZnJlcXBlbl9jb250cm9sbGVkIikNCg0KbV9oY19tYXNmcmVxX2NvbnRyb2xsZWQgPSBicm0oZGlhcnlfbWFzdHVyYmF0aW9uX3N1bSB+DQogICAgICAgICAgICAgICAgICAgICAgICBvZmZzZXQobG9nKG51bWJlcl9vZl9kYXlzKSkgKw0KICAgICAgICAgICAgICAgICAgICAgICAgY29udHJhY2VwdGlvbl9ob3Jtb25hbCArDQogICAgICAgICAgICAgICAgICAgICAgICBhZ2UgKyBuZXRfaW5jb21lICsgcmVsYXRpb25zaGlwX2R1cmF0aW9uX2ZhY3RvciArDQogICAgICAgICAgICAgICAgICAgICAgICAgICAgICBlZHVjYXRpb25feWVhcnMgKw0KICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgYmZpX2V4dHJhICsgYmZpX25ldXJvICsgYmZpX2FncmVlICsgYmZpX2NvbnNjICsgYmZpX29wZW4gKw0KICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgcmVsaWdpb3NpdHksDQogICAgICAgICAgICAgICAgZGF0YSA9IGRhdGEsIGZhbWlseSA9IHBvaXNzb24oKSwNCiAgICAgICAgICAgICAgICBmaWxlID0gIm1yb2J1c3RfaGNfbWFzZnJlcV9jb250cm9sbGVkIikNCmBgYA0KDQojIyMjIEZvcmVzdCBQbG90IGZvciBFZmZlY3QgU2l6ZXMgey5hY3RpdmV9DQpgYGB7cn0NCm1vZGVscyA9IHJiaW5kKA0KICBtX2hjX2F0cnJfY29udHJvbGxlZCAlPiUNCiAgICBzcHJlYWRfZHJhd3MoYl9jb250cmFjZXB0aW9uX2hvcm1vbmFseWVzKSAlPiUNCiAgICBwaXZvdF9sb25nZXIoY29scyA9IGMoYl9jb250cmFjZXB0aW9uX2hvcm1vbmFseWVzKSwNCiAgICAgICAgICAgICAgICAgbmFtZXNfdG8gPSAiY29uZGl0aW9uIiwNCiAgICAgICAgICAgICAgICAgdmFsdWVzX3RvID0gInJfY29uZGl0aW9uIikgJT4lDQogICAgbXV0YXRlKG1vZGVsID0gIlBlcmNlaXZlZCBQYXJ0bmVyIEF0dHJhY3RpdmVuZXNzXG5ST1BFID0gWy0wLjA3LCAwLjA3XSIsDQogICAgICAgICAgIFJPUEUgPSAwLjA3KSwNCiAgbV9oY19yZWxzYXRfY29udHJvbGxlZCAlPiUNCiAgICBzcHJlYWRfZHJhd3MoYl9jb250cmFjZXB0aW9uX2hvcm1vbmFseWVzKSAlPiUNCiAgICBwaXZvdF9sb25nZXIoY29scyA9IGMoYl9jb250cmFjZXB0aW9uX2hvcm1vbmFseWVzKSwNCiAgICAgICAgICAgICAgICAgbmFtZXNfdG8gPSAiY29uZGl0aW9uIiwNCiAgICAgICAgICAgICAgICAgdmFsdWVzX3RvID0gInJfY29uZGl0aW9uIikgJT4lDQogICAgbXV0YXRlKG1vZGVsID0gIlJlbGF0aW9uc2hpcCBTYXRpc2ZhY3Rpb25cblJPUEUgPSBbLTAuMDQsIDAuMDRdIiwNCiAgICAgICAgICAgUk9QRSA9IDAuMDQpLA0KICBtX2hjX3NleHNhdF9jb250cm9sbGVkICU+JQ0KICAgIHNwcmVhZF9kcmF3cyhiX2NvbnRyYWNlcHRpb25faG9ybW9uYWx5ZXMpICU+JQ0KICAgIHBpdm90X2xvbmdlcihjb2xzID0gYyhiX2NvbnRyYWNlcHRpb25faG9ybW9uYWx5ZXMpLA0KICAgICAgICAgICAgICAgICBuYW1lc190byA9ICJjb25kaXRpb24iLA0KICAgICAgICAgICAgICAgICB2YWx1ZXNfdG8gPSAicl9jb25kaXRpb24iKSAlPiUNCiAgICBtdXRhdGUobW9kZWwgPSAiU2V4dWFsIFNhdGlzZmFjdGlvblxuUk9QRSA9IFstMC4xMSwgMC4xMV0iLA0KICAgICAgICAgICBST1BFID0gMC4xMSksDQogIG1faGNfbGliaWRvX2NvbnRyb2xsZWQgJT4lDQogICAgc3ByZWFkX2RyYXdzKGJfY29udHJhY2VwdGlvbl9ob3Jtb25hbHllcykgJT4lDQogICAgcGl2b3RfbG9uZ2VyKGNvbHMgPSBjKGJfY29udHJhY2VwdGlvbl9ob3Jtb25hbHllcyksDQogICAgICAgICAgICAgICAgIG5hbWVzX3RvID0gImNvbmRpdGlvbiIsDQogICAgICAgICAgICAgICAgIHZhbHVlc190byA9ICJyX2NvbmRpdGlvbiIpICU+JQ0KICAgIG11dGF0ZShtb2RlbCA9ICJMaWJpZG9cblJPUEUgPSBbLTAuMDYsIDAuMDZdIiwNCiAgICAgICAgICAgUk9QRSA9IDAuMDYpLA0KICBtX2hjX3NleGZyZXFwZW5fY29udHJvbGxlZCAlPiUNCiAgICBzcHJlYWRfZHJhd3MoYl9jb250cmFjZXB0aW9uX2hvcm1vbmFseWVzKSAlPiUNCiAgICBwaXZvdF9sb25nZXIoY29scyA9IGMoYl9jb250cmFjZXB0aW9uX2hvcm1vbmFseWVzKSwNCiAgICAgICAgICAgICAgICAgbmFtZXNfdG8gPSAiY29uZGl0aW9uIiwNCiAgICAgICAgICAgICAgICAgdmFsdWVzX3RvID0gInJfY29uZGl0aW9uIikgJT4lDQogICAgbXV0YXRlKG1vZGVsID0gIkZyZXF1ZW5jeSBvZiBWYWdpbmFsIEludGVyY291cnNlXG5ST1BFID0gWy0wLjA1LCAwLjA1XSIsDQogICAgICAgICAgIFJPUEUgPSAwLjA1KSwNCiAgbV9oY19tYXNmcmVxX2NvbnRyb2xsZWQgJT4lDQogICAgc3ByZWFkX2RyYXdzKGJfY29udHJhY2VwdGlvbl9ob3Jtb25hbHllcykgJT4lDQogICAgcGl2b3RfbG9uZ2VyKGNvbHMgPSBjKGJfY29udHJhY2VwdGlvbl9ob3Jtb25hbHllcyksDQogICAgICAgICAgICAgICAgIG5hbWVzX3RvID0gImNvbmRpdGlvbiIsDQogICAgICAgICAgICAgICAgIHZhbHVlc190byA9ICJyX2NvbmRpdGlvbiIpICU+JQ0KICAgIG11dGF0ZShtb2RlbCA9ICJGcmVxdWVuY3kgb2YgTWFzdHVyYmF0aW9uXG5ST1BFID0gWy0wLjA1LCAwLjA1XSIsDQogICAgICAgICAgIFJPUEUgPSAwLjA1KSkNCg0KZGF0YTEgPSBtb2RlbHMgJT4lDQogIG11dGF0ZShtb2RlbCA9IGZhY3Rvcihtb2RlbCwNCiAgICAgICAgICAgICAgICAgICBsZXZlbHMgPSBjKCJGcmVxdWVuY3kgb2YgTWFzdHVyYmF0aW9uXG5ST1BFID0gWy0wLjA1LCAwLjA1XSIsDQogICAgICAgICAgICAgICAgICAgICAgICAgICAgICAiRnJlcXVlbmN5IG9mIFZhZ2luYWwgSW50ZXJjb3Vyc2VcblJPUEUgPSBbLTAuMDUsIDAuMDVdIiwNCiAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICJMaWJpZG9cblJPUEUgPSBbLTAuMDYsIDAuMDZdIiwNCiAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICJTZXh1YWwgU2F0aXNmYWN0aW9uXG5ST1BFID0gWy0wLjExLCAwLjExXSIsDQogICAgICAgICAgICAgICAgICAgICAgICAgICAgICAiUmVsYXRpb25zaGlwIFNhdGlzZmFjdGlvblxuUk9QRSA9IFstMC4wNCwgMC4wNF0iLA0KICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgIlBlcmNlaXZlZCBQYXJ0bmVyIEF0dHJhY3RpdmVuZXNzXG5ST1BFID0gWy0wLjA3LCAwLjA3XSIpKSkNCmRhdGEyID0gbW9kZWxzICU+JQ0KICBncm91cF9ieShtb2RlbCwgUk9QRSkgJT4lDQogIG11dGF0ZShtb2RlbCA9IGZhY3Rvcihtb2RlbCwNCiAgICAgICAgICAgICAgICAgICAgICAgIGxldmVscyA9IGMoIkZyZXF1ZW5jeSBvZiBNYXN0dXJiYXRpb25cblJPUEUgPSBbLTAuMDUsIDAuMDVdIiwNCiAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgIkZyZXF1ZW5jeSBvZiBWYWdpbmFsIEludGVyY291cnNlXG5ST1BFID0gWy0wLjA1LCAwLjA1XSIsDQogICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICJMaWJpZG9cblJPUEUgPSBbLTAuMDYsIDAuMDZdIiwNCiAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgIlNleHVhbCBTYXRpc2ZhY3Rpb25cblJPUEUgPSBbLTAuMTEsIDAuMTFdIiwNCiAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgIlJlbGF0aW9uc2hpcCBTYXRpc2ZhY3Rpb25cblJPUEUgPSBbLTAuMDQsIDAuMDRdIiwNCiAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgIlBlcmNlaXZlZCBQYXJ0bmVyIEF0dHJhY3RpdmVuZXNzXG5ST1BFID0gWy0wLjA3LCAwLjA3XSIpKSkgJT4lDQogIG1lYW5fcWkoY29uZGl0aW9uX21lYW4gPSByX2NvbmRpdGlvbiwNCiAgICAgICAgICAud2lkdGggPSBjKC45MCkpICU+JQ0KICBtdXRhdGUodGV4dCA9IGlmZWxzZSgud2lkdGggPT0gLjkwLA0KICAgICAgICAgICAgICAgICAgICAgICBzdHJfYyhyb3VuZChjb25kaXRpb25fbWVhbiwgMiksDQogICAgICAgICAgICAgICAgICAgICAgICAgICAgICIgWyIsDQogICAgICAgICAgICAgICAgICAgICAgICAgICAgIHJvdW5kKC5sb3dlciwgMiksDQogICAgICAgICAgICAgICAgICAgICAgICAgICAgICI7ICIsDQogICAgICAgICAgICAgICAgICAgICAgICAgICAgIHJvdW5kKC51cHBlciwgMiksDQogICAgICAgICAgICAgICAgICAgICAgICAgICAgICJdIiksIE5BKSwNCiAgICAgICAgIHNpZyA9IGlmZWxzZSgubG93ZXIgPiBST1BFIHwgLnVwcGVyIDwgLVJPUEUsICJkZWNsaW5lZCIsDQogICAgICAgICAgICAgICAgICAgICAgaWZlbHNlKGFicygubG93ZXIpIDwgUk9QRSAgJiAudXBwZXIgPiBST1BFLCAidW5kZWNpZGFibGUiLCAiYWNjZXB0ZWQiKSkpDQoNCg0KaGNfY29udHJvbGxlZCA9IGdncGxvdCgpICsNCiAgc3RhdF9oYWxmZXllKGRhdGEgPSBkYXRhMSwNCiAgICAgICAgICAgICAgIGFlcyh5ID0gbW9kZWwsIHggPSByX2NvbmRpdGlvbiwNCiAgICAgICAgICAgICAgICAgICBjb2xvciA9ICJncmV5IiksDQogICAgICAgICAgICAgICAud2lkdGggPSAuOTApICsNCiAgZ2VvbV92bGluZSh4aW50ZXJjZXB0ID0gMCwgbGluZXR5cGUgPSAiZG90dGVkIikgKw0KICBnZW9tX3BvaW50aW50ZXJ2YWwoZGF0YSA9IGRhdGEyLA0KICAgICAgICAgICAgICAgICAgICAgYWVzKHkgPSBtb2RlbCwNCiAgICAgICAgICAgICAgICAgICAgICAgICB4ID0gY29uZGl0aW9uX21lYW4sDQogICAgICAgICAgICAgICAgICAgICAgICAgeG1pbiA9IC5sb3dlciwNCiAgICAgICAgICAgICAgICAgICAgICAgICB4bWF4ID0gLnVwcGVyLA0KICAgICAgICAgICAgICAgICAgICAgICAgIGNvbG9yID0gc2lnKSkgKw0KICBnZW9tX3BvaW50KGRhdGEgPSBkYXRhMSwgYWVzKHkgPSBtb2RlbCwgeCA9IFJPUEUpLCBzaGFwZSA9ICJ8Iiwgc2l6ZSA9IDEwLCBjb2xvciA9ICJibGFjayIpICsNCiAgZ2VvbV9wb2ludChkYXRhID0gZGF0YTEsIGFlcyh5ID0gbW9kZWwsIHggPSAtUk9QRSksIHNoYXBlID0gInwiLCBzaXplID0gMTAsIGNvbG9yID0gImJsYWNrIikgKw0KICBnZW9tX3BvaW50KGRhdGEgPSBkYXRhMSwgYWVzKHkgPSBtb2RlbCwgeCA9IFJPUEUpLCBzaGFwZSA9IDU4LCBzaXplID0gMTAsIGNvbG9yID0gIndoaXRlIikgKw0KICBnZW9tX3BvaW50KGRhdGEgPSBkYXRhMSwgYWVzKHkgPSBtb2RlbCwgeCA9IC1ST1BFKSwgc2hhcGUgPSA1OCwgc2l6ZSA9IDEwLCBjb2xvciA9ICJ3aGl0ZSIpICsNCiAgYXBhdGhlbWUgKw0KICB0aGVtZShsZWdlbmQucG9zaXRpb24gPSAibm9uZSIpICsNCiAgc2NhbGVfY29sb3JfbWFudWFsKHZhbHVlcyA9IGMoInJlZCIsICJncmVlbiIsICJza3libHVlIiwgImdyZXkiKSkgKw0KICBzY2FsZV94X2NvbnRpbnVvdXMoYnJlYWtzID0gYygtMC41LCAtMC40LCAtMC4zLCAtMC4yLCAtMC4xLCAwLCAwLjEsIDAuMiwgMC4zLCAwLjQsIDAuNSksDQogICAgICAgICAgICAgICAgICAgICBsaW1pdHMgPSBjKC0wLjUsIDAuNSkpICsNCiAgbGFicyh4ID0gIkVmZmVjdCBTaXplIEVzdGltYXRlcyBvZiBIb3Jtb25hbCBDb250cmFjZXB0aW9uIiwgeSA9ICJPdXRjb21lcyIpICArDQogIHRoZW1lKGxlZ2VuZC5wb3NpdGlvbiA9ICJub25lIikNCmhjX2NvbnRyb2xsZWQNCmBgYA0KDQojIyMgQ29tYmluZSBQbG90cw0KYGBge3J9DQpoY19wbG90cyA9DQogIGdnYXJyYW5nZShoY191bmNvbnRyb2xsZWQgKyB0aGVtZShwbG90Lm1hcmdpbiA9IG1hcmdpbigxLDAsMCwwLCJjbSIpKSwNCiAgICAgICAgICAgIGhjX2NvbnRyb2xsZWQgKyB0aGVtZShwbG90Lm1hcmdpbiA9IG1hcmdpbigzLDAsMCwwLCJjbSIpKSwNCiAgICAgICAgICBsYWJlbHMgPSBjKCJVbmNvbnRyb2xsZWQgTW9kZWwiLCAiXG5Db250cm9sbGVkIE1vZGVsIiksDQogICAgICAgICAgZm9udC5sYWJlbCA9IGxpc3Qoc2l6ZSA9IDIwLCBjb2xvciA9ICJibGFjayIsIGZhY2UgPSAiYm9sZCIsIGZhbWlseSA9ICJzYW5zIiksDQogICAgICAgICAgaGp1c3QgPSAtMC4xLA0KICAgICAgICAgIG5jb2wgPSAxLCBucm93ID0gMiwNCiAgICAgICAgICBhbGlnbiA9ICJ2IikNCg0KaGNfcGxvdHMNCmpwZWcoJ0VmZmVjdCBzaXplIGVzdGltYXRlcyBmb3IgaG9ybW9uYWwgY29udHJhY2VwdGl2ZXMgb24gb3V0Y29tZXMuanBnJywgDQogICAgIHdpZHRoID0gODAwLCBoZWlnaHQgPSAxMDAwLCBxdWFsaXR5ID0gMTAwKQ0KaGNfcGxvdHMNCmRldi5vZmYoKQ0KYGBgDQoNCiMjIEVmZmVjdHMgb2YgKGluKWNvbmdydWVudCB1c2Ugb2YgaG9ybW9uYWwgY29udHJhY2VwdGl2ZXMNCiMjIyBQbG90cw0KYGBge3J9DQptX2NvbmdydWVuY3lfYXRyciA9IGJybShhdHRyYWN0aXZlbmVzc19wYXJ0bmVyIH4NCiAgICAgICAgICAgICAgICAgICAgICAgICAgY29udHJhY2VwdGlvbl9ob3Jtb25hbCAqIGNvbmdydWVudF9jb250cmFjZXB0aW9uLA0KICAgICAgICAgICAgICAgIGRhdGEgPSBkYXRhLCBmYW1pbHkgPSBnYXVzc2lhbigpLA0KICAgICAgICAgICAgICAgIGZpbGUgPSAibV9jb25ncnVlbmN5X2F0cnIiKQ0KDQptX2NvbmdydWVuY3lfYXRycl9jZSA9IG1fY29uZ3J1ZW5jeV9hdHJyICU+JSANCiAgY29uZGl0aW9uYWxfZWZmZWN0cyguLCBlZmZlY3RzID0gImNvbnRyYWNlcHRpb25faG9ybW9uYWw6Y29uZ3J1ZW50X2NvbnRyYWNlcHRpb24iLA0KICAgICAgICAgICAgICAgICAgICBwcm9iID0gMC45MCkNCg0KbV9jb25ncnVlbmN5X2F0cnJfcGxvdCA9IA0KICBwbG90KG1fY29uZ3J1ZW5jeV9hdHJyX2NlLCBwbG90ID0gRkFMU0UpW1sxXV0gKw0KICBsYWJzKHggPSAiQ3VycmVudCBDb250cmFjZXB0aXZlIE1ldGhvZCIsDQogICAgICAgeSA9ICJQZXJjZWl2ZWQgUGFydG5lciBBdHRyYWN0aXZlbmVzcyIpICsNCiAgc2NhbGVfeF9kaXNjcmV0ZShsYWJlbHM9YygiTm9uLUhDIiwgIkhDIikpICsNCiAgc2NhbGVfY29sb3JfbWFudWFsKG5hbWUgPSAiQ29uZ3J1ZW50IENvbnRyYWNlcHRpdmUgTWV0aG9kIiwgbGFiZWxzID0gYygiTm8iLCAiWWVzIiksDQogICAgICAgICAgICAgICAgICAgICB2YWx1ZXM9YygiI0Q5NUYwMiIsICIjMUI5RTc3IikpICsNCiAgc2NhbGVfZmlsbF9tYW51YWwobmFtZSA9ICJDb25ncnVlbnQgQ29udHJhY2VwdGl2ZSBNZXRob2QiLCBsYWJlbHMgPSBjKCJObyIsICJZZXMiKSwNCiAgICAgICAgICAgICAgICAgICAgdmFsdWVzPWMoIiNEOTVGMDIiLCAiIzFCOUU3NyIpKSArDQogIGFwYXRoZW1lDQoNCm1fY29uZ3J1ZW5jeV9hdHJyX3Bsb3QNCg0KIG1fY29uZ3J1ZW5jeV9yZWxzYXQgPSBicm0ocmVsYXRpb25zaGlwX3NhdGlzZmFjdGlvbiB+DQogICAgICAgICAgICAgICAgICAgICAgICAgIGNvbnRyYWNlcHRpb25faG9ybW9uYWwgKiBjb25ncnVlbnRfY29udHJhY2VwdGlvbiwNCiAgICAgICAgICAgICAgICBkYXRhID0gZGF0YSwgZmFtaWx5ID0gZ2F1c3NpYW4oKSwNCiAgICAgICAgICAgICAgICBmaWxlID0gIm1fY29uZ3J1ZW5jeV9yZWxzYXQiKQ0KDQptX2NvbmdydWVuY3lfcmVsc2F0X2NlID0gbV9jb25ncnVlbmN5X3JlbHNhdCAlPiUgDQogIGNvbmRpdGlvbmFsX2VmZmVjdHMoLiwgZWZmZWN0cyA9ICJjb250cmFjZXB0aW9uX2hvcm1vbmFsOmNvbmdydWVudF9jb250cmFjZXB0aW9uIiwNCiAgICAgICAgICAgICAgICAgICAgcHJvYiA9IDAuOTApDQoNCm1fY29uZ3J1ZW5jeV9yZWxzYXRfcGxvdCA9IA0KICBwbG90KG1fY29uZ3J1ZW5jeV9yZWxzYXRfY2UsIHBsb3QgPSBGQUxTRSlbWzFdXSArDQogIGxhYnMoeCA9ICJDdXJyZW50IENvbnRyYWNlcHRpdmUgTWV0aG9kIiwNCiAgICAgICB5ID0gIlJlbGF0aW9uc2hpcCBTYXRpc2ZhY3Rpb24iKSArDQogIHNjYWxlX3hfZGlzY3JldGUobGFiZWxzPWMoIk5vbi1IQyIsICJIQyIpKSArDQogIHNjYWxlX2NvbG9yX21hbnVhbChuYW1lID0gIkNvbmdydWVudCBDb250cmFjZXB0aXZlIE1ldGhvZCIsIGxhYmVscyA9IGMoIk5vIiwgIlllcyIpLA0KICAgICAgICAgICAgICAgICAgICAgdmFsdWVzPWMoIiNEOTVGMDIiLCAiIzFCOUU3NyIpKSArDQogIHNjYWxlX2ZpbGxfbWFudWFsKG5hbWUgPSAiQ29uZ3J1ZW50IENvbnRyYWNlcHRpdmUgTWV0aG9kIiwgbGFiZWxzID0gYygiTm8iLCAiWWVzIiksDQogICAgICAgICAgICAgICAgICAgIHZhbHVlcz1jKCIjRDk1RjAyIiwgIiMxQjlFNzciKSkgKw0KICBhcGF0aGVtZQ0KDQptX2NvbmdydWVuY3lfcmVsc2F0X3Bsb3QNCg0KbV9jb25ncnVlbmN5X3NleHNhdCA9IGJybShzYXRpc2ZhY3Rpb25fc2V4dWFsX2ludGVyY291cnNlIH4NCiAgICAgICAgICAgICAgICAgICAgICAgICAgY29udHJhY2VwdGlvbl9ob3Jtb25hbCAqIGNvbmdydWVudF9jb250cmFjZXB0aW9uLA0KICAgICAgICAgICAgICAgIGRhdGEgPSBkYXRhLCBmYW1pbHkgPSBnYXVzc2lhbigpLA0KICAgICAgICAgICAgICAgIGZpbGUgPSAibV9jb25ncnVlbmN5X3NleHNhdCIpDQoNCg0KbV9jb25ncnVlbmN5X3NleHNhdF9jZSA9IG1fY29uZ3J1ZW5jeV9zZXhzYXQgJT4lIA0KICBjb25kaXRpb25hbF9lZmZlY3RzKC4sIGVmZmVjdHMgPSAiY29udHJhY2VwdGlvbl9ob3Jtb25hbDpjb25ncnVlbnRfY29udHJhY2VwdGlvbiIsDQogICAgICAgICAgICAgICAgICAgIHByb2IgPSAwLjkwKQ0KDQptX2NvbmdydWVuY3lfc2V4c2F0X3Bsb3QgPSANCiAgcGxvdChtX2NvbmdydWVuY3lfc2V4c2F0X2NlLCBwbG90ID0gRkFMU0UpW1sxXV0gKw0KICBsYWJzKHggPSAiQ3VycmVudCBDb250cmFjZXB0aXZlIE1ldGhvZCIsDQogICAgICAgeSA9ICJTZXh1YWwgU2F0aXNmYWN0aW9uIikgKw0KICBzY2FsZV94X2Rpc2NyZXRlKGxhYmVscz1jKCJOb24tSEMiLCAiSEMiKSkgKw0KICBzY2FsZV9jb2xvcl9tYW51YWwobmFtZSA9ICJDb25ncnVlbnQgQ29udHJhY2VwdGl2ZSBNZXRob2QiLCBsYWJlbHMgPSBjKCJObyIsICJZZXMiKSwNCiAgICAgICAgICAgICAgICAgICAgIHZhbHVlcz1jKCIjRDk1RjAyIiwgIiMxQjlFNzciKSkgKw0KICBzY2FsZV9maWxsX21hbnVhbChuYW1lID0gIkNvbmdydWVudCBDb250cmFjZXB0aXZlIE1ldGhvZCIsIGxhYmVscyA9IGMoIk5vIiwgIlllcyIpLA0KICAgICAgICAgICAgICAgICAgICB2YWx1ZXM9YygiI0Q5NUYwMiIsICIjMUI5RTc3IikpICsNCiAgYXBhdGhlbWUNCg0KbV9jb25ncnVlbmN5X3NleHNhdF9wbG90DQoNCm1fY29uZ3J1ZW5jeV9saWJpZG8gPSBicm0oZGlhcnlfbGliaWRvX21lYW4gfg0KICAgICAgICAgICAgICAgICAgICAgICAgICBjb250cmFjZXB0aW9uX2hvcm1vbmFsICogY29uZ3J1ZW50X2NvbnRyYWNlcHRpb24sDQogICAgICAgICAgICAgICAgZGF0YSA9IGRhdGEsIGZhbWlseSA9IGdhdXNzaWFuKCksDQogICAgICAgICAgICAgICAgZmlsZSA9ICJtX2NvbmdydWVuY3lfbGliaWRvIikNCg0KbV9jb25ncnVlbmN5X2xpYmlkb19jZSA9IG1fY29uZ3J1ZW5jeV9saWJpZG8gJT4lIA0KICBjb25kaXRpb25hbF9lZmZlY3RzKC4sIGVmZmVjdHMgPSAiY29udHJhY2VwdGlvbl9ob3Jtb25hbDpjb25ncnVlbnRfY29udHJhY2VwdGlvbiIsDQogICAgICAgICAgICAgICAgICAgIHByb2IgPSAwLjkwKQ0KDQptX2NvbmdydWVuY3lfbGliaWRvX3Bsb3QgPSANCiAgcGxvdChtX2NvbmdydWVuY3lfbGliaWRvX2NlLCBwbG90ID0gRkFMU0UpW1sxXV0gKw0KICBsYWJzKHggPSAiQ3VycmVudCBDb250cmFjZXB0aXZlIE1ldGhvZCIsDQogICAgICAgeSA9ICJMaWJpZG8iKSArDQogIHNjYWxlX3hfZGlzY3JldGUobGFiZWxzPWMoIk5vbi1IQyIsICJIQyIpKSArDQogIHNjYWxlX2NvbG9yX21hbnVhbChuYW1lID0gIkNvbmdydWVudCBDb250cmFjZXB0aXZlIE1ldGhvZCIsIGxhYmVscyA9IGMoIk5vIiwgIlllcyIpLA0KICAgICAgICAgICAgICAgICAgICAgdmFsdWVzPWMoIiNEOTVGMDIiLCAiIzFCOUU3NyIpKSArDQogIHNjYWxlX2ZpbGxfbWFudWFsKG5hbWUgPSAiQ29uZ3J1ZW50IENvbnRyYWNlcHRpdmUgTWV0aG9kIiwgbGFiZWxzID0gYygiTm8iLCAiWWVzIiksDQogICAgICAgICAgICAgICAgICAgIHZhbHVlcz1jKCIjRDk1RjAyIiwgIiMxQjlFNzciKSkgKw0KICBhcGF0aGVtZQ0KDQptX2NvbmdydWVuY3lfbGliaWRvX3Bsb3QNCg0KbV9jb25ncnVlbmN5X3NleGZyZXFwZW4gPSBicm0oZGlhcnlfc2V4X2FjdGl2ZV9zZXhfc3VtIH4gb2Zmc2V0KGxvZyhudW1iZXJfb2ZfZGF5cykpICsNCiAgICAgICAgICAgICAgICAgICAgIGNvbnRyYWNlcHRpb25faG9ybW9uYWwgKiBjb25ncnVlbnRfY29udHJhY2VwdGlvbiwNCiAgICAgICAgICAgICAgICBkYXRhID0gZGF0YSwgZmFtaWx5ID0gcG9pc3NvbigpLA0KICAgICAgICAgICAgICAgIGZpbGUgPSAibXJvYnVzdF9jb25ncnVlbmN5X3NleGZyZXFwZW4iKQ0KDQoNCm1fY29uZ3J1ZW5jeV9zZXhmcmVxcGVuX2NlID0gbV9jb25ncnVlbmN5X3NleGZyZXFwZW4gJT4lIA0KICBjb25kaXRpb25hbF9lZmZlY3RzKC4sIGVmZmVjdHMgPSAiY29udHJhY2VwdGlvbl9ob3Jtb25hbDpjb25ncnVlbnRfY29udHJhY2VwdGlvbiIsDQogICAgICAgICAgICAgICAgICAgIHByb2IgPSAwLjkwLA0KICAgICAgICAgICAgICAgICAgICBjb25kaXRpb25zID0gZGF0YS5mcmFtZShudW1iZXJfb2ZfZGF5cyA9IDEpKQ0KDQptX2NvbmdydWVuY3lfc2V4ZnJlcXBlbl9wbG90ID0gDQogIHBsb3QobV9jb25ncnVlbmN5X3NleGZyZXFwZW5fY2UsIHBsb3QgPSBGQUxTRSlbWzFdXSArDQogIGxhYnMoeCA9ICJDdXJyZW50IENvbnRyYWNlcHRpdmUgTWV0aG9kIiwNCiAgICAgICB5ID0gIkZyZXF1ZW5jeSBvZiBWYWdpbmFsIEludGVyY291cnNlXG4oUHJvYmFibGl0eSBwZXIgRGF5KSIpICsNCiAgc2NhbGVfeF9kaXNjcmV0ZShsYWJlbHM9YygiTm9uLUhDIiwgIkhDIikpICsNCiAgc2NhbGVfY29sb3JfbWFudWFsKG5hbWUgPSAiQ29uZ3J1ZW50IENvbnRyYWNlcHRpdmUgTWV0aG9kIiwgbGFiZWxzID0gYygiTm8iLCAiWWVzIiksDQogICAgICAgICAgICAgICAgICAgICB2YWx1ZXM9YygiI0Q5NUYwMiIsICIjMUI5RTc3IikpICsNCiAgc2NhbGVfZmlsbF9tYW51YWwobmFtZSA9ICJDb25ncnVlbnQgQ29udHJhY2VwdGl2ZSBNZXRob2QiLCBsYWJlbHMgPSBjKCJObyIsICJZZXMiKSwNCiAgICAgICAgICAgICAgICAgICAgdmFsdWVzPWMoIiNEOTVGMDIiLCAiIzFCOUU3NyIpKSArDQogIGFwYXRoZW1lIA0KDQptX2NvbmdydWVuY3lfc2V4ZnJlcXBlbl9wbG90DQoNCm1fY29uZ3J1ZW5jeV9tYXNmcmVxID0gYnJtKGRpYXJ5X21hc3R1cmJhdGlvbl9zdW0gfiBvZmZzZXQobG9nKG51bWJlcl9vZl9kYXlzKSkgKw0KICAgICAgICAgICAgICAgICAgICAgICAgICAgICBjb250cmFjZXB0aW9uX2hvcm1vbmFsICogY29uZ3J1ZW50X2NvbnRyYWNlcHRpb24sDQogICAgICAgICAgICAgICAgZGF0YSA9IGRhdGEsIGZhbWlseSA9IHBvaXNzb24oKSwNCiAgICAgICAgICAgICAgICBmaWxlID0gIm1yb2J1c3RfY29uZ3J1ZW5jeV9tYXNmcmVxIikNCg0KbV9jb25ncnVlbmN5X21hc2ZyZXFfY2UgPSBtX2NvbmdydWVuY3lfbWFzZnJlcSAlPiUgDQogIGNvbmRpdGlvbmFsX2VmZmVjdHMoLiwgZWZmZWN0cyA9ICJjb250cmFjZXB0aW9uX2hvcm1vbmFsOmNvbmdydWVudF9jb250cmFjZXB0aW9uIiwNCiAgICAgICAgICAgICAgICAgICAgcHJvYiA9IDAuOTAsDQogICAgICAgICAgICAgICAgICAgIGNvbmRpdGlvbnMgPSBkYXRhLmZyYW1lKG51bWJlcl9vZl9kYXlzID0gMSkpDQoNCm1fY29uZ3J1ZW5jeV9tYXNmcmVxX3Bsb3QgPSANCiAgcGxvdChtX2NvbmdydWVuY3lfbWFzZnJlcV9jZSwgcGxvdCA9IEZBTFNFKVtbMV1dICsNCiAgbGFicyh4ID0gIkN1cnJlbnQgQ29udHJhY2VwdGl2ZSBNZXRob2QiLA0KICAgICAgIHkgPSAiRnJlcXVlbmN5IG9mIE1hc3R1cmJhdGlvblxuKFByb2JhYmxpdHkgcGVyIERheSkiKSArDQogIHNjYWxlX3hfZGlzY3JldGUobGFiZWxzPWMoIk5vbi1IQyIsICJIQyIpKSArDQogIHNjYWxlX2NvbG9yX21hbnVhbChuYW1lID0gIkNvbmdydWVudCBDb250cmFjZXB0aXZlIE1ldGhvZCIsIGxhYmVscyA9IGMoIk5vIiwgIlllcyIpLA0KICAgICAgICAgICAgICAgICAgICAgdmFsdWVzPWMoIiNEOTVGMDIiLCAiIzFCOUU3NyIpKSArDQogIHNjYWxlX2ZpbGxfbWFudWFsKG5hbWUgPSAiQ29uZ3J1ZW50IENvbnRyYWNlcHRpdmUgTWV0aG9kIiwgbGFiZWxzID0gYygiTm8iLCAiWWVzIiksDQogICAgICAgICAgICAgICAgICAgIHZhbHVlcz1jKCIjRDk1RjAyIiwgIiMxQjlFNzciKSkgKw0KICBhcGF0aGVtZQ0KDQptX2NvbmdydWVuY3lfbWFzZnJlcV9wbG90DQpgYGANCg0KIyMjIENvbWJpbmUgUGxvdHMNCmBgYHtyfQ0KY29uZ3J1ZW5jeV91bmNvbnRyb2xsZWQgPQ0KICBnZ2FycmFuZ2UobV9jb25ncnVlbmN5X2F0cnJfcGxvdCArDQogICAgICAgICAgICAgIHJyZW1vdmUoInhsYWIiKSwNCiAgICAgICAgICAgIG1fY29uZ3J1ZW5jeV9yZWxzYXRfcGxvdCArDQogICAgICAgICAgICAgIHJyZW1vdmUoInhsYWIiKSwNCiAgICAgICAgICAgIG1fY29uZ3J1ZW5jeV9zZXhzYXRfcGxvdCArDQogICAgICAgICAgICAgIHJyZW1vdmUoInhsYWIiKSwNCiAgICAgICAgICAgIG1fY29uZ3J1ZW5jeV9saWJpZG9fcGxvdCArDQogICAgICAgICAgICAgIHJyZW1vdmUoInhsYWIiKSwNCiAgICAgICAgICAgIG1fY29uZ3J1ZW5jeV9zZXhmcmVxcGVuX3Bsb3QsDQogICAgICAgICAgICBtX2NvbmdydWVuY3lfbWFzZnJlcV9wbG90LA0KICAgICAgICAgIGNvbW1vbi5sZWdlbmQgPSBUUlVFLCBsZWdlbmQgPSAiYm90dG9tIiwNCiAgICAgICAgICBuY29sID0gMiwgbnJvdyA9IDMsDQogICAgICAgICAgYWxpZ24gPSAidiIpDQoNCmNvbmdydWVuY3lfdW5jb250cm9sbGVkDQpqcGVnKCdEaWZmZXJlbmNlcyBpbiBjdXJyZW50IGNvbnRyYWNlcHRpdmUgbWV0aG9kLCBjb25ncnVlbnQgdXNlIG9mIGNvbnRyYWNlcHRpdmUgbWV0aG9kLCBhbmQgdGhlaXIgaW50ZXJhY3Rpb24uanBnJywgDQogICAgIHdpZHRoID0gODAwLCBoZWlnaHQgPSAxMDAwLCBxdWFsaXR5ID0gMTAwKQ0KY29uZ3J1ZW5jeV91bmNvbnRyb2xsZWQNCmRldi5vZmYoKQ0KYGBgDQoNCiMjIENvbnRyb2xsZWQgZWZmZWN0cyBvZiAoaW4pY29uZ3J1ZW50IHVzZSBvZiBob3Jtb25hbCBjb250cmFjZXB0aXZlcw0KIyMjIFBsb3RzDQpgYGB7cn0NCm1fY29uZ3J1ZW5jeV9hdHJyX2NvbnRyb2xsZWQgPSBicm0oYXR0cmFjdGl2ZW5lc3NfcGFydG5lciB+DQogICAgICAgICAgICAgICAgICAgICAgICAgIGNvbnRyYWNlcHRpb25faG9ybW9uYWwgKiBjb25ncnVlbnRfY29udHJhY2VwdGlvbiArDQogICAgICAgICAgICAgICAgICAgICAgICAgICAgIGFnZSArIG5ldF9pbmNvbWUgKyByZWxhdGlvbnNoaXBfZHVyYXRpb25fZmFjdG9yICsNCiAgICAgICAgICAgICAgICAgICAgICAgICAgICAgIGVkdWNhdGlvbl95ZWFycyArDQogICAgICAgICAgICAgICAgICAgICAgICAgICAgICBiZmlfZXh0cmEgKyBiZmlfbmV1cm8gKyBiZmlfYWdyZWUgKyBiZmlfY29uc2MgKyBiZmlfb3BlbiArDQogICAgICAgICAgICAgICAgICAgICAgICAgICAgICByZWxpZ2lvc2l0eSwNCiAgICAgICAgICAgICAgICBkYXRhID0gZGF0YSwgZmFtaWx5ID0gZ2F1c3NpYW4oKSwNCiAgICAgICAgICAgICAgICBmaWxlID0gIm1fY29uZ3J1ZW5jeV9hdHJyX2NvbnRyb2xsZWQiKQ0KDQptX2NvbmdydWVuY3lfYXRycl9jb250cm9sbGVkX2NlID0gbV9jb25ncnVlbmN5X2F0cnJfY29udHJvbGxlZCAlPiUgDQogIGNvbmRpdGlvbmFsX2VmZmVjdHMoLiwgZWZmZWN0cyA9ICJjb250cmFjZXB0aW9uX2hvcm1vbmFsOmNvbmdydWVudF9jb250cmFjZXB0aW9uIiwNCiAgICAgICAgICAgICAgICAgICAgcHJvYiA9IDAuOTApDQoNCm1fY29uZ3J1ZW5jeV9hdHJyX2NvbnRyb2xsZWRfcGxvdCA9IA0KICBwbG90KG1fY29uZ3J1ZW5jeV9hdHJyX2NvbnRyb2xsZWRfY2UsIHBsb3QgPSBGQUxTRSlbWzFdXSArDQogIGxhYnMoeCA9ICJDdXJyZW50IENvbnRyYWNlcHRpdmUgTWV0aG9kIiwNCiAgICAgICB5ID0gIlBlcmNlaXZlZCBQYXJ0bmVyIEF0dHJhY3RpdmVuZXNzIikgKw0KICBzY2FsZV94X2Rpc2NyZXRlKGxhYmVscz1jKCJOb24tSEMiLCAiSEMiKSkgKw0KICBzY2FsZV9jb2xvcl9tYW51YWwobmFtZSA9ICJDb25ncnVlbnQgQ29udHJhY2VwdGl2ZSBNZXRob2QiLCBsYWJlbHMgPSBjKCJObyIsICJZZXMiKSwNCiAgICAgICAgICAgICAgICAgICAgIHZhbHVlcz1jKCIjRDk1RjAyIiwgIiMxQjlFNzciKSkgKw0KICBzY2FsZV9maWxsX21hbnVhbChuYW1lID0gIkNvbmdydWVudCBDb250cmFjZXB0aXZlIE1ldGhvZCIsIGxhYmVscyA9IGMoIk5vIiwgIlllcyIpLA0KICAgICAgICAgICAgICAgICAgICB2YWx1ZXM9YygiI0Q5NUYwMiIsICIjMUI5RTc3IikpICsNCiAgYXBhdGhlbWUNCg0KbV9jb25ncnVlbmN5X2F0cnJfY29udHJvbGxlZF9wbG90DQoNCm1fY29uZ3J1ZW5jeV9yZWxzYXRfY29udHJvbGxlZCA9IGJybShyZWxhdGlvbnNoaXBfc2F0aXNmYWN0aW9uIH4NCiAgICAgICAgICAgICAgICAgICAgICAgICAgY29udHJhY2VwdGlvbl9ob3Jtb25hbCAqIGNvbmdydWVudF9jb250cmFjZXB0aW9uICsNCiAgICAgICAgICAgICAgICAgICAgICAgICAgICAgYWdlICsgbmV0X2luY29tZSArIHJlbGF0aW9uc2hpcF9kdXJhdGlvbl9mYWN0b3IgKw0KICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgZWR1Y2F0aW9uX3llYXJzICsNCiAgICAgICAgICAgICAgICAgICAgICAgICAgICAgIGJmaV9leHRyYSArIGJmaV9uZXVybyArIGJmaV9hZ3JlZSArIGJmaV9jb25zYyArIGJmaV9vcGVuICsNCiAgICAgICAgICAgICAgICAgICAgICAgICAgICAgIHJlbGlnaW9zaXR5LA0KICAgICAgICAgICAgICAgIGRhdGEgPSBkYXRhLCBmYW1pbHkgPSBnYXVzc2lhbigpLA0KICAgICAgICAgICAgICAgIGZpbGUgPSAibV9jb25ncnVlbmN5X3JlbHNhdF9jb250cm9sbGVkIikNCg0KbV9jb25ncnVlbmN5X3JlbHNhdF9jb250cm9sbGVkX2NlID0gbV9jb25ncnVlbmN5X3JlbHNhdF9jb250cm9sbGVkICU+JSANCiAgY29uZGl0aW9uYWxfZWZmZWN0cyguLCBlZmZlY3RzID0gImNvbnRyYWNlcHRpb25faG9ybW9uYWw6Y29uZ3J1ZW50X2NvbnRyYWNlcHRpb24iLA0KICAgICAgICAgICAgICAgICAgICBwcm9iID0gMC45MCkNCg0KbV9jb25ncnVlbmN5X3JlbHNhdF9jb250cm9sbGVkX3Bsb3QgPSANCiAgcGxvdChtX2NvbmdydWVuY3lfcmVsc2F0X2NvbnRyb2xsZWRfY2UsIHBsb3QgPSBGQUxTRSlbWzFdXSArDQogIGxhYnMoeCA9ICJDdXJyZW50IENvbnRyYWNlcHRpdmUgTWV0aG9kIiwNCiAgICAgICB5ID0gIlJlbGF0aW9uc2hpcCBTYXRpc2ZhY3Rpb24iKSArDQogIHNjYWxlX3hfZGlzY3JldGUobGFiZWxzPWMoIk5vbi1IQyIsICJIQyIpKSArDQogIHNjYWxlX2NvbG9yX21hbnVhbChuYW1lID0gIkNvbmdydWVudCBDb250cmFjZXB0aXZlIE1ldGhvZCIsIGxhYmVscyA9IGMoIk5vIiwgIlllcyIpLA0KICAgICAgICAgICAgICAgICAgICAgdmFsdWVzPWMoIiNEOTVGMDIiLCAiIzFCOUU3NyIpKSArDQogIHNjYWxlX2ZpbGxfbWFudWFsKG5hbWUgPSAiQ29uZ3J1ZW50IENvbnRyYWNlcHRpdmUgTWV0aG9kIiwgbGFiZWxzID0gYygiTm8iLCAiWWVzIiksDQogICAgICAgICAgICAgICAgICAgIHZhbHVlcz1jKCIjRDk1RjAyIiwgIiMxQjlFNzciKSkgKw0KICBhcGF0aGVtZSANCg0KbV9jb25ncnVlbmN5X3JlbHNhdF9jb250cm9sbGVkX3Bsb3QNCg0KbV9jb25ncnVlbmN5X3NleHNhdF9jb250cm9sbGVkID0gYnJtKHNhdGlzZmFjdGlvbl9zZXh1YWxfaW50ZXJjb3Vyc2Ugfg0KICAgICAgICAgICAgICAgICAgICAgICAgICBjb250cmFjZXB0aW9uX2hvcm1vbmFsICogY29uZ3J1ZW50X2NvbnRyYWNlcHRpb24gKw0KICAgICAgICAgICAgICAgICAgICAgICAgICAgICBhZ2UgKyBuZXRfaW5jb21lICsgcmVsYXRpb25zaGlwX2R1cmF0aW9uX2ZhY3RvciArDQogICAgICAgICAgICAgICAgICAgICAgICAgICAgICBlZHVjYXRpb25feWVhcnMgKw0KICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgYmZpX2V4dHJhICsgYmZpX25ldXJvICsgYmZpX2FncmVlICsgYmZpX2NvbnNjICsgYmZpX29wZW4gKw0KICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgcmVsaWdpb3NpdHksDQogICAgICAgICAgICAgICAgZGF0YSA9IGRhdGEsIGZhbWlseSA9IGdhdXNzaWFuKCksDQogICAgICAgICAgICAgICAgZmlsZSA9ICJtX2NvbmdydWVuY3lfc2V4c2F0X2NvbnRyb2xsZWQiKQ0KDQptX2NvbmdydWVuY3lfc2V4c2F0X2NvbnRyb2xsZWRfY2UgPSBtX2NvbmdydWVuY3lfc2V4c2F0X2NvbnRyb2xsZWQgJT4lIA0KICBjb25kaXRpb25hbF9lZmZlY3RzKC4sIGVmZmVjdHMgPSAiY29udHJhY2VwdGlvbl9ob3Jtb25hbDpjb25ncnVlbnRfY29udHJhY2VwdGlvbiIsDQogICAgICAgICAgICAgICAgICAgIHByb2IgPSAwLjkwKQ0KDQptX2NvbmdydWVuY3lfc2V4c2F0X2NvbnRyb2xsZWRfcGxvdCA9IA0KICBwbG90KG1fY29uZ3J1ZW5jeV9zZXhzYXRfY29udHJvbGxlZF9jZSwgcGxvdCA9IEZBTFNFKVtbMV1dICsNCiAgbGFicyh4ID0gIkN1cnJlbnQgQ29udHJhY2VwdGl2ZSBNZXRob2QiLA0KICAgICAgIHkgPSAiU2V4dWFsIFNhdGlzZmFjdGlvbiIpICsNCiAgc2NhbGVfeF9kaXNjcmV0ZShsYWJlbHM9YygiTm9uLUhDIiwgIkhDIikpICsNCiAgc2NhbGVfY29sb3JfbWFudWFsKG5hbWUgPSAiQ29uZ3J1ZW50IENvbnRyYWNlcHRpdmUgTWV0aG9kIiwgbGFiZWxzID0gYygiTm8iLCAiWWVzIiksDQogICAgICAgICAgICAgICAgICAgICB2YWx1ZXM9YygiI0Q5NUYwMiIsICIjMUI5RTc3IikpICsNCiAgc2NhbGVfZmlsbF9tYW51YWwobmFtZSA9ICJDb25ncnVlbnQgQ29udHJhY2VwdGl2ZSBNZXRob2QiLCBsYWJlbHMgPSBjKCJObyIsICJZZXMiKSwNCiAgICAgICAgICAgICAgICAgICAgdmFsdWVzPWMoIiNEOTVGMDIiLCAiIzFCOUU3NyIpKSArDQogIGFwYXRoZW1lIA0KDQptX2NvbmdydWVuY3lfc2V4c2F0X2NvbnRyb2xsZWRfcGxvdA0KDQptX2NvbmdydWVuY3lfbGliaWRvX2NvbnRyb2xsZWQgPSBicm0oZGlhcnlfbGliaWRvX21lYW4gfg0KICAgICAgICAgICAgICAgICAgICAgICAgICBjb250cmFjZXB0aW9uX2hvcm1vbmFsICogY29uZ3J1ZW50X2NvbnRyYWNlcHRpb24gKw0KICAgICAgICAgICAgICAgICAgICAgICAgICAgICBhZ2UgKyBuZXRfaW5jb21lICsgcmVsYXRpb25zaGlwX2R1cmF0aW9uX2ZhY3RvciArDQogICAgICAgICAgICAgICAgICAgICAgICAgICAgICBlZHVjYXRpb25feWVhcnMgKw0KICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgYmZpX2V4dHJhICsgYmZpX25ldXJvICsgYmZpX2FncmVlICsgYmZpX2NvbnNjICsgYmZpX29wZW4gKw0KICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgcmVsaWdpb3NpdHksDQogICAgICAgICAgICAgICAgZGF0YSA9IGRhdGEsIGZhbWlseSA9IGdhdXNzaWFuKCksDQogICAgICAgICAgICAgICAgZmlsZSA9ICJtX2NvbmdydWVuY3lfbGliaWRvX2NvbnRyb2xsZWQiKQ0KDQptX2NvbmdydWVuY3lfbGliaWRvX2NvbnRyb2xsZWRfY2UgPSBtX2NvbmdydWVuY3lfbGliaWRvX2NvbnRyb2xsZWQgJT4lIA0KICBjb25kaXRpb25hbF9lZmZlY3RzKC4sIGVmZmVjdHMgPSAiY29udHJhY2VwdGlvbl9ob3Jtb25hbDpjb25ncnVlbnRfY29udHJhY2VwdGlvbiIsDQogICAgICAgICAgICAgICAgICAgIHByb2IgPSAwLjkwKQ0KDQptX2NvbmdydWVuY3lfbGliaWRvX2NvbnRyb2xsZWRfcGxvdCA9IA0KICBwbG90KG1fY29uZ3J1ZW5jeV9saWJpZG9fY29udHJvbGxlZF9jZSwgcGxvdCA9IEZBTFNFKVtbMV1dICsNCiAgbGFicyh4ID0gIkN1cnJlbnQgQ29udHJhY2VwdGl2ZSBNZXRob2QiLA0KICAgICAgIHkgPSAiTGliaWRvIikgKw0KICBzY2FsZV94X2Rpc2NyZXRlKGxhYmVscz1jKCJOb24tSEMiLCAiSEMiKSkgKw0KICBzY2FsZV9jb2xvcl9tYW51YWwobmFtZSA9ICJDb25ncnVlbnQgQ29udHJhY2VwdGl2ZSBNZXRob2QiLCBsYWJlbHMgPSBjKCJObyIsICJZZXMiKSwNCiAgICAgICAgICAgICAgICAgICAgIHZhbHVlcz1jKCIjRDk1RjAyIiwgIiMxQjlFNzciKSkgKw0KICBzY2FsZV9maWxsX21hbnVhbChuYW1lID0gIkNvbmdydWVudCBDb250cmFjZXB0aXZlIE1ldGhvZCIsIGxhYmVscyA9IGMoIk5vIiwgIlllcyIpLA0KICAgICAgICAgICAgICAgICAgICB2YWx1ZXM9YygiI0Q5NUYwMiIsICIjMUI5RTc3IikpICsNCiAgYXBhdGhlbWUgDQoNCm1fY29uZ3J1ZW5jeV9saWJpZG9fY29udHJvbGxlZF9wbG90DQoNCm1fY29uZ3J1ZW5jeV9zZXhmcmVxcGVuX2NvbnRyb2xsZWQgPSBicm0oZGlhcnlfc2V4X2FjdGl2ZV9zZXhfc3VtIH4NCiAgICAgICAgICAgICAgICAgICAgICAgIG9mZnNldChsb2cobnVtYmVyX29mX2RheXMpKSArDQogICAgICAgICAgICAgICAgICAgICAgICBjb250cmFjZXB0aW9uX2hvcm1vbmFsICogY29uZ3J1ZW50X2NvbnRyYWNlcHRpb24gKw0KICAgICAgICAgICAgICAgICAgICAgICAgYWdlICsgbmV0X2luY29tZSArIHJlbGF0aW9uc2hpcF9kdXJhdGlvbl9mYWN0b3IgKw0KICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgZWR1Y2F0aW9uX3llYXJzICsNCiAgICAgICAgICAgICAgICAgICAgICAgICAgICAgIGJmaV9leHRyYSArIGJmaV9uZXVybyArIGJmaV9hZ3JlZSArIGJmaV9jb25zYyArIGJmaV9vcGVuICsNCiAgICAgICAgICAgICAgICAgICAgICAgICAgICAgIHJlbGlnaW9zaXR5LA0KICAgICAgICAgICAgICAgIGRhdGEgPSBkYXRhLCBmYW1pbHkgPSBwb2lzc29uKCksDQogICAgICAgICAgICAgICAgZmlsZSA9ICJtcm9idXN0X2NvbmdydWVuY3lfc2V4ZnJlcXBlbl9jb250cm9sbGVkIikNCg0KbV9jb25ncnVlbmN5X3NleGZyZXFwZW5fY29udHJvbGxlZF9jZSA9IG1fY29uZ3J1ZW5jeV9zZXhmcmVxcGVuX2NvbnRyb2xsZWQgJT4lIA0KICBjb25kaXRpb25hbF9lZmZlY3RzKC4sIGVmZmVjdHMgPSAiY29udHJhY2VwdGlvbl9ob3Jtb25hbDpjb25ncnVlbnRfY29udHJhY2VwdGlvbiIsDQogICAgICAgICAgICAgICAgICAgIHByb2IgPSAwLjkwLA0KICAgICAgICAgICAgICAgICAgICBjb25kaXRpb25zID0gZGF0YS5mcmFtZShudW1iZXJfb2ZfZGF5cyA9IDEpKQ0KDQptX2NvbmdydWVuY3lfc2V4ZnJlcXBlbl9jb250cm9sbGVkX3Bsb3QgPSANCiAgcGxvdChtX2NvbmdydWVuY3lfc2V4ZnJlcXBlbl9jb250cm9sbGVkX2NlLCBwbG90ID0gRkFMU0UpW1sxXV0gKw0KICBsYWJzKHggPSAiQ3VycmVudCBDb250cmFjZXB0aXZlIE1ldGhvZCIsDQogICAgICAgeSA9ICJGcmVxdWVuY3kgb2YgVmFnaW5hbCBJbnRlcmNvdXJzZVxuKFByb2JhYmlsaXR5IHBlciBEYXkpIikgKw0KICBzY2FsZV94X2Rpc2NyZXRlKGxhYmVscz1jKCJOb24tSEMiLCAiSEMiKSkgKw0KICBzY2FsZV9jb2xvcl9tYW51YWwobmFtZSA9ICJDb25ncnVlbnQgQ29udHJhY2VwdGl2ZSBNZXRob2QiLCBsYWJlbHMgPSBjKCJObyIsICJZZXMiKSwNCiAgICAgICAgICAgICAgICAgICAgIHZhbHVlcz1jKCIjRDk1RjAyIiwgIiMxQjlFNzciKSkgKw0KICBzY2FsZV9maWxsX21hbnVhbChuYW1lID0gIkNvbmdydWVudCBDb250cmFjZXB0aXZlIE1ldGhvZCIsIGxhYmVscyA9IGMoIk5vIiwgIlllcyIpLA0KICAgICAgICAgICAgICAgICAgICB2YWx1ZXM9YygiI0Q5NUYwMiIsICIjMUI5RTc3IikpICsNCiAgYXBhdGhlbWUgDQoNCm1fY29uZ3J1ZW5jeV9zZXhmcmVxcGVuX2NvbnRyb2xsZWRfcGxvdA0KDQptX2NvbmdydWVuY3lfbWFzZnJlcV9jb250cm9sbGVkID0gYnJtKGRpYXJ5X21hc3R1cmJhdGlvbl9zdW0gfg0KICAgICAgICAgICAgICAgICAgICAgICAgb2Zmc2V0KGxvZyhudW1iZXJfb2ZfZGF5cykpICsNCiAgICAgICAgICAgICAgICAgICAgICAgIGNvbnRyYWNlcHRpb25faG9ybW9uYWwgKiBjb25ncnVlbnRfY29udHJhY2VwdGlvbiArDQogICAgICAgICAgICAgICAgICAgICAgICBhZ2UgKyBuZXRfaW5jb21lICsgcmVsYXRpb25zaGlwX2R1cmF0aW9uX2ZhY3RvciArDQogICAgICAgICAgICAgICAgICAgICAgICAgICAgICBlZHVjYXRpb25feWVhcnMgKw0KICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgYmZpX2V4dHJhICsgYmZpX25ldXJvICsgYmZpX2FncmVlICsgYmZpX2NvbnNjICsgYmZpX29wZW4gKw0KICAgICAgICAgICAgICAgICAgICAgICAgICAgICAgcmVsaWdpb3NpdHksDQogICAgICAgICAgICAgICAgZGF0YSA9IGRhdGEsIGZhbWlseSA9IHBvaXNzb24oKSwNCiAgICAgICAgICAgICAgICBmaWxlID0gIm1yb2J1c3RfY29uZ3J1ZW5jeV9tYXNmcmVxX2NvbnRyb2xsZWQiKQ0KDQptX2NvbmdydWVuY3lfbWFzZnJlcV9jb250cm9sbGVkX2NlID0gbV9jb25ncnVlbmN5X21hc2ZyZXFfY29udHJvbGxlZCAlPiUgDQogIGNvbmRpdGlvbmFsX2VmZmVjdHMoLiwgZWZmZWN0cyA9ICJjb250cmFjZXB0aW9uX2hvcm1vbmFsOmNvbmdydWVudF9jb250cmFjZXB0aW9uIiwNCiAgICAgICAgICAgICAgICAgICAgcHJvYiA9IDAuOTAsDQogICAgICAgICAgICAgICAgICAgIGNvbmRpdGlvbnMgPSBkYXRhLmZyYW1lKG51bWJlcl9vZl9kYXlzID0gMSkpDQoNCm1fY29uZ3J1ZW5jeV9tYXNmcmVxX2NvbnRyb2xsZWRfcGxvdCA9IA0KICBwbG90KG1fY29uZ3J1ZW5jeV9tYXNmcmVxX2NvbnRyb2xsZWRfY2UsIHBsb3QgPSBGQUxTRSlbWzFdXSArDQogIGxhYnMoeCA9ICJDdXJyZW50IENvbnRyYWNlcHRpdmUgTWV0aG9kIiwNCiAgICAgICB5ID0gIkZyZXF1ZW5jeSBvZiBNYXN0dXJiYXRpb25cbihQcm9iYWJpbGl0eSBwZXIgRGF5KSIpICsNCiAgc2NhbGVfeF9kaXNjcmV0ZShsYWJlbHM9YygiTm9uLUhDIiwgIkhDIikpICsNCiAgc2NhbGVfY29sb3JfbWFudWFsKG5hbWUgPSAiQ29uZ3J1ZW50IENvbnRyYWNlcHRpdmUgTWV0aG9kIiwgbGFiZWxzID0gYygiTm8iLCAiWWVzIiksDQogICAgICAgICAgICAgICAgICAgICB2YWx1ZXM9YygiI0Q5NUYwMiIsICIjMUI5RTc3IikpICsNCiAgc2NhbGVfZmlsbF9tYW51YWwobmFtZSA9ICJDb25ncnVlbnQgQ29udHJhY2VwdGl2ZSBNZXRob2QiLCBsYWJlbHMgPSBjKCJObyIsICJZZXMiKSwNCiAgICAgICAgICAgICAgICAgICAgdmFsdWVzPWMoIiNEOTVGMDIiLCAiIzFCOUU3NyIpKSArDQogIGFwYXRoZW1lIA0KDQptX2NvbmdydWVuY3lfbWFzZnJlcV9jb250cm9sbGVkX3Bsb3QNCmBgYA0KDQojIyMgQ29tYmluZSBQbG90cw0KYGBge3J9DQpjb25ncnVlbmN5X2NvbnRyb2xsZWQgPQ0KICBnZ2FycmFuZ2UobV9jb25ncnVlbmN5X2F0cnJfY29udHJvbGxlZF9wbG90ICsgcnJlbW92ZSgieGxhYiIpLA0KICAgICAgICAgICAgbV9jb25ncnVlbmN5X3JlbHNhdF9jb250cm9sbGVkX3Bsb3QgKyBycmVtb3ZlKCJ4bGFiIiksDQogICAgICAgICAgbV9jb25ncnVlbmN5X3NleHNhdF9jb250cm9sbGVkX3Bsb3QgKyBycmVtb3ZlKCJ4bGFiIiksDQogICAgICAgICAgbV9jb25ncnVlbmN5X2xpYmlkb19jb250cm9sbGVkX3Bsb3QgKyBycmVtb3ZlKCJ4bGFiIiksDQogICAgICAgICAgbV9jb25ncnVlbmN5X3NleGZyZXFwZW5fY29udHJvbGxlZF9wbG90LA0KICAgICAgICAgIG1fY29uZ3J1ZW5jeV9tYXNmcmVxX2NvbnRyb2xsZWRfcGxvdCwNCiAgICAgICAgICBjb21tb24ubGVnZW5kID0gVFJVRSwgbGVnZW5kID0gImJvdHRvbSIsDQogICAgICAgICAgbmNvbCA9IDIsIG5yb3cgPSAzLA0KICAgICAgICAgIGFsaWduID0gInYiKQ0KDQpjb25ncnVlbmN5X2NvbnRyb2xsZWQNCmpwZWcoJ0RpZmZlcmVuY2VzIGluIGN1cnJlbnQgY29udHJhY2VwdGl2ZSBtZXRob2QsIGNvbmdydWVudCB1c2Ugb2YgY29udHJhY2VwdGl2ZSBtZXRob2QsIGFuZCB0aGVpciBpbnRlcmFjdGlvbiBpbiBjb250cm9sbGVkIG1vZGVscy5qcGcnLCANCiAgICAgd2lkdGggPSA4MDAsIGhlaWdodCA9IDEwMDAsIHF1YWxpdHkgPSAxMDApDQpjb25ncnVlbmN5X2NvbnRyb2xsZWQNCmRldi5vZmYoKQ0KYGBgDQoNCg==